|
 |
| Korean J Intern Med > Volume 36(Suppl 1); 2021 > Article |
|
Abstract
Background/Aims
Methods
Results
Conclusions
KEY MESSAGE
Acknowledgments
Supplementary Materials
Supplementary Table 1.
Supplementary Table 2.
Supplementary Table 3.
Supplementary Table 4.
Figure 1.
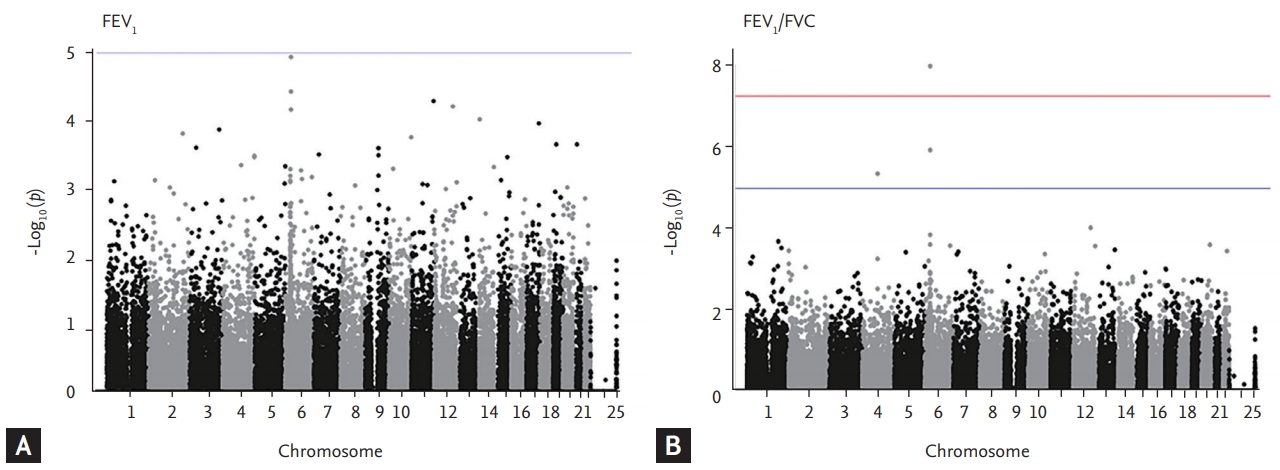
Figure 2.

Table 1.
Table 2.
| Trait |
KoGES |
UK biobank |
CHARGE consortium |
SpiroMeta consortium |
||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SNP ID | Chr:Pos | Gene | MAF | OR or beta (QT) | p value | OR or beta (QT) | p value | OR or beta (QT) | p value | OR or beta (QT) | p value | |
| FEV1 | rs7742369 | 6:34,165,721 | - | 0.171 | 3.75E-02 | 7.11 × 10-5 | 0.028 | 4.00E-31 | - | - | - | - |
| rs2780226 | 6:34,199,092 | - | 0.165 | 3.94E-02 | 3.86 × 10-5 | –0.035 | 1.15E-26 | - | - | - | - | |
| rs1150781 | 6:34,214,322 | C6orf1 (SMIM29) | 0.181 | 4.04E-02 | 1.20 × 10-5 | –0.033 | 5.65E-24 | - | - | - | - | |
| 11:130,247,700 | 11:130,247,700 | - | 0.464 | –2.82E-02 | 5.36 × 10-5 | - | - | - | - | - | - | |
| 12:110,390,979 | 12:110,390,979 | GIT2/TCHP | 0.078 | –5.31E-02 | 6.55 × 10-5 | - | - | - | - | - | - | |
| rs114591848 | 14:21,550,212 | ARHGEF40 | 0.014 | 1.17E-01 | 9.95 × 10-5 | - | - | - | - | - | - | |
| rs7143633a | 14:21,547,088 | ARHGEF40 | - | - | - | 2.33 | 2.36E-03 | - | - | - | - | |
| FEV1/FVC | rs7671167 | 4:89,883,979 | FAM13A | 0.493 | 4.74E-01 | 4.54 × 10-6 | - | - | 0.307 | 3.60E-09 | 0.043 | 5.87E-06 |
| rs2045517 | 4:89,870,964 | FAM13A | - | - | - | –0.047 | 2.00E-11 | 0.438 | 1.60E-8 | –0.047 | 2.00E-11 | |
| rs2239688 | 6:32,054,212 | TNXB | 0.175 | 7.03E-01 | 1.30 × 10-6 | - | - | - | - | - | - | |
| rs433061b | 6:32,054,212 | TNXB | - | - | - | - | - | 0.127 | 2.00E-04 | - | - | |
| rs1150754c | 6:32,050,758 | TNXB | - | - | - | - | - | 0.111 | 3.00E-03 | - | - | |
| rs2070600 | 6:32,151,443 | AGER | 0.158 | 8.75E-01 | 1.09 × 10-8 | 0.126 | 9.07E-15 | 0.186 | 1.50E-08 | - | 2.56E-02 | |
Total 31,571 SNPs for FEV1 and total 16,616 SNPs for FEV1/FVC analyzed. Results presented for the trait and chromosome (Chr) and position (Pos) in build 37 are given for each SNP.
FEV1, forced expiratory volume in 1 second; FVC, forced vital capacity; KoGES, The Korean Genome Epidemiology Study; CHARGE, Cohorts for Heart and Aging Research in Genomic Epidemiology; SNP, single nucleotide polymorphism; Chr, chromosome; Pos, position; MAF, minor allele frequency; OR, odds ratio; SMIM29, small integral membrane protein 29; GIT2, G protein-coupled receptor kinase interacting ArfGAP2; TCHP, trichoplein keratin filament binding; ARHGEF40, rho guanine nucleotide exchange factor 40; FAM13A, family with sequence similarity 13 member A; TNXB, tenascin XB; AGER, advanced glycosylation end-product specific receptor.
Table 3.
Results are given as chromosome, trait and p values (p < 2.5 × 10-5).
SKAT, Sequence Kernel Association Test; KoGES, Korean Genome and Epidemiology Study; Chr, chromosome; DFFA, DNA fragmentation factor subunit alpha; FEV1, forced expiratory volume in 1 second; CHST15, carbohydrate sulfotransferase 15; FVC, forced vital capacity; DEFB135, defensin beta 135; MOGAT2, monoacylglycerol O-acyltransferase 2; BSX, brain specific homeobox; SPTY2D1, SPT2 chromatin protein domain containing 1; DEFB118, defensin beta 118; PTGDR, prostaglandin D2 receptor; MYBL2, MYB proto-oncogene like 2.



 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Download Citation
Download Citation Supplement 1
Supplement 1 Print
Print