|
 |
| Korean J Intern Med > Volume 24(1); 2009 > Article |
|
Abstract
Background/Aims
Mutations of the epidermal growth factor receptor (EGFR) and Kirsten rat sarcoma viral oncogene (KRAS) are important in the pathogenesis of lung cancer, and recent reports have revealed racial and geographical differences in mutation expression.
Methods
This study was conducted to investigate the prevalence of EGFR and KRAS mutations and their correlation with clinical variables in Korean patients with adenocarcinoma of the lung. Formalin-fixed adenocarcinoma specimens from 104 randomly selected patients diagnosed at Kosin University Gospel Hospital from October 1996 to January 2005 were used for the study.
Results
We found a high prevalence of EGFR mutations and a low prevalence of KRAS mutations. EGFR mutations were present in 24% (25 of 104) of the samples: one mutation in exon 18, 13 in exon 19, one in exon 20, and 10 in exon 21. The presence of an EGFR mutation was not associated with gender, smoking history, histological grade, age, bronchioalveolar components, or cancer stage in patients with adenocarcinoma of the lung.
The human genome encodes approximately 518 kinases, including 90 tyrosine kinases (TK) and 43 tyrosine-like kinases [1]. Phosphorylated tyrosine serves as a binding site for several signal transducers that initiate multiple signaling pathways, resulting in cell proliferation, migration, metastasis, resistance to apoptosis, and angiogenesis [2]. Epidermal growth factor receptor (EGFR) appears to play a central role in tumorigenesis, and targeting this receptor may provide a unique approach for treating EGFR-expressing cancers [3]. This concept led to the development and regulatory approval of small-molecule EGFR TK inhibitors (TKIs) such as gefitinib and erlotinib; however, clinical trials have revealed significant variability in the response to TKIs. Good clinical responses have been observed most frequently in women, nonsmokers, patients with adenocarcinomas, and patients of East Asian origin [4,5]. Activating mutations of EGFR have been reported in a subset of patients with pulmonary adenocarcinomas, and tumors with EGFR mutations are highly sensitive to gefitinib [6,7]. Furthermore, the incidence of EGFR mutations is higher in East Asians compared with Caucasian patients and in those with adenocarcinoma [7].
Therefore, we investigated the presence of EGFR mutations in randomly selected Korean patients with adenocarcinoma of the lung, to determine the correlation between clinical features and the presence of Kirsten rat sarcoma viral oncogene (KRAS) mutations.
Primary tumor samples from patients with adenocarcinoma of the lung were obtained from 104 randomly selected patients with lung cancer who were diagnosed at Kosin University Gospel Hospital, Busan, Korea from October 1996 to January 2005. Clinical information, including gender, age at diagnosis, and smoking history, were obtained by a retrospective review of the medical history recorded in the patients' charts. We excluded patients with metastatic adenocarcinoma and those who were treated with gefitinib, because treatment with TKIs alters the prognosis of adenocarcinoma in East Asian patients. The stage of the disease was the postoperative pathological stage in patients who had undergone surgery and the pretreated stage for non-surgically treated patients.
DNA was extracted from five paraffin-embedded sections of 10-┬ĄM thickness representing a portion of each tumor block, using a QIAamp DNA Mini kit (Qiagen, Hilden, Germany). EGFR exons were amplified from the DNA (100 ng) by polymerase chain reaction (PCR) in a 20-┬ĄL solution containing 2 ┬ĄL of 10├Śbuffer (Roche, Mannheim, Germany), 1.7 to 2.5 mmol/L of MgCl2, 0.3 ┬ĄM of each EGFR primer pair (exon 18, F: 5'-tccaaatgagctggcaagtg, R: 5'-tcccaaacactcagtgaaacaaa; exon 19, F: 5'-atgtggcaccatctcacaattgcc, R: 5'-ccacacagcaaagcagaaactcac; exon 20, F:5'-cattcatgcgtcttcacctg, R: 5'-catatccccatggcaaactc; exon 21, F: 5'-gctcagagcctggcatgaa, R: 5'-cat cctcccctgcatgtgt), 250 ┬Ąmol/L of deoxynucleotide triphosphates, and 2.5 units of DNA polymerase (Roche). Amplifications were performed using a 5-min initial denaturation at 94Ōäā followed by 30 cycles of 1 min at 94Ōäā, 1 min at 57Ōäā, and 1 min at 72Ōäā, with a final 10-min extension at 72Ōäā. KRAS was similarly amplified from the DNA, except that a primer pair for KRAS codons 12 and 13 was used (F: 5'-ttatgtgtgacatgttctaat and R: 5'-agaatggtcctgcaccagtaa) and the amplification conditions were a 5-min initial denaturation at 94Ōäā; 30 cycles of 1 min at 94Ōäā, and 1 min at 55Ōäā, 1 min at 72Ōäā; and a final 10-min extension at 72Ōäā. All PCR products were purified from a 2% agarose gel, using a QIAgen gel extraction kit (Qiagen), and used as templates for DNA sequencing with both forward and reverse sequence-specific primers.
For sequencing, each purified PCR product (20 ng) was used in a 20-┬ĄL sequencing reaction solution containing 8 ┬ĄL of BigDye Terminator v3.1 (Applied Biosystems, Foster City, CA) and 0.1 ┬Ąmol/L of the appropriate PCR primers. Sequencing was performed with 25 cycles of 10 s at 96Ōäā, 5 s at 50Ōäā, and 4 min at 60Ōäā, using an ABI PRISM 3100 DNA Analyzer (Applied Biosystems). The data were analyzed using Sequencer 3.1.1. software (Applied Biosystems), to compare the sequence variations.
The statistical analyses of categorical variables were performed using Pearson's Žć2 test or Fisher's exact test, as appropriate. The median duration for overall patient survival was calculated using the Kaplan-Meier method. Comparisons between groups were made using the logrank test. Multivariate analysis was carried out using the stepwise Cox regression model. Two-sided p values <0.05 were considered significant. All analyses were performed using SPSS for Windows, version 12.0 (SPSS Inc., Chicago, IL).
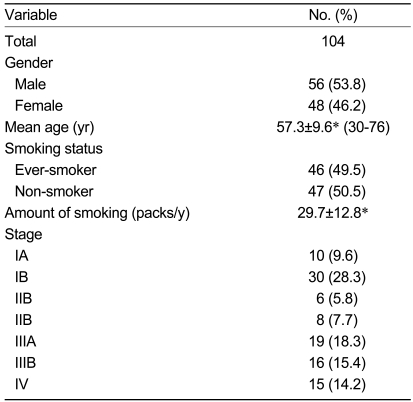
There were 104 patients (male, 56; female, 48), with a mean age of 57.3 years (range, 30-76). Forty-six patients were current or ex-smokers. The cancer stages of the patients are shown in Table 1. Eighty-one (77.9%) patients underwent surgery, 17 patients received chemotherapy with or without radiotherapy, and six patients received only supportive care.
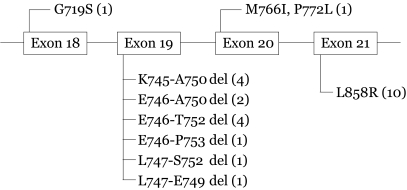
A mutation of the EGFR gene was detected in 25 (24%) of the 104 tumors: one mutation in exon 18 (G719S point mutation); 13 in exon 19 (9-18 bp in- frame deletion); one in exon 20 (M766L/P772L point mutation); and 10 in exon 21 (L858R point mutation) (Fig. 1). There were no tumors with multiple mutations. In exon 19, all 13 of the mutations were small in-frame deletions of the kinase domain; four were K745-A750del, two were E746-A750del, four were E746-T753del, one was E746-P753del, one was L747-S752, and one was L747-E749del. In exon 21, all 10 mutations were a substitution of T for G at the second nucleotide of codon 858, resulting in the substitution of an arginine for a leucine in the protein. The mutation in exon 18 was a substitution of A for G at nucleotide 2155, producing a glycine instead of a serine at codon 719. The mutation in exon 20 substituted G for A and C for T at nucleotide 2298 and resulted in an amino acid substitution of methionine for isoleucine at codon 766 and proline for leucine at codon 772.
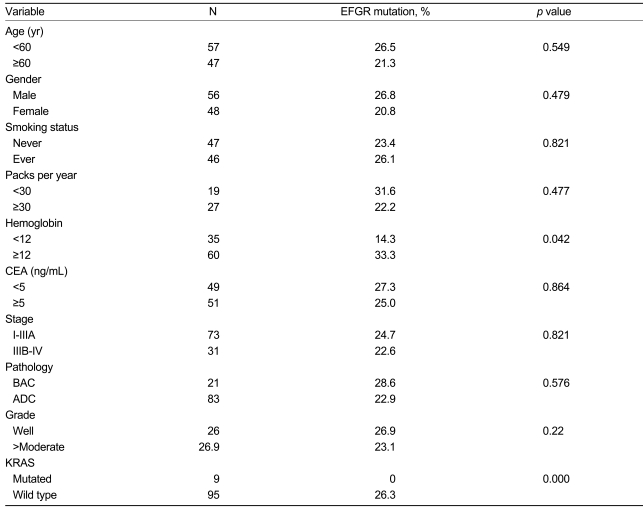
The presence of EGFR mutations was not associated with age, clinical stage, gender, smoking history, or carcinoembryonic antigen (CEA) level. There was also no correlation between EGFR mutations and the pathological subtype or differentiation in these patients. However, EGFR mutations were more frequently associated with a high hemoglobin level (14.3 vs. 33.3%, <12 g/dL vs. Ōēź12 g/dL; p=0.042) (Table 2).
KRAS mutations were present in 9.6% [9] of 94 tumors (8 mutations in codon 12 and 1 in codon 13). EGFR mutations were never found in tumors with KRAS mutations, suggesting a mutually exclusive relationship.
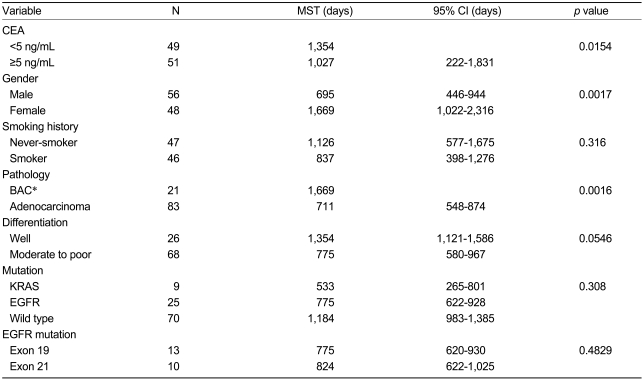
At the time of the analysis, 49 deaths (47.1%) had occurred. The Kaplan-Meier curves indicated that the site of the EGFR mutation did not affect patient prognosis (median survival time: 775 days with exon 19 mutation vs. 824 days with exon 21 mutation; p=0.483), and smoking history was not associated with survival outcome. The survival time for patients with KRAS mutations tended to be shorter than that for patients with EGFR mutations (median survival time: 17.5 vs. 25.5 months), but the difference was not statistically significant.
Table 3 shows the results of the univariate analysis. Female patients had a longer median survival time than male patients (p=0.0017), and patients with a higher CEA serum level had a longer median survival time than the other patients (p=0.0154). Patients with bronchioloalveolar cell carcinoma and alveolar cell components had a longer median survival time than other adenocarcinoma patients (p=0.0016). The median survival time of the well-differentiated carcinoma patients was longer than that of patients with moderate to poorly differentiated carcinoma (p=0.0546). According to the univariate analysis, stage, CEA level, gender, smoking history, pathological subtype, and differentiation were the covariates selected for potential inclusion in a multivariate Cox regression model of patient survival. The selected model included stage (p=0.0000), gender (p=0.027), pathological subtype (p=0.003), and differentiation (p=0.024) (Table 4).
Adenocarcinoma is the most common histological subtype of lung cancer, and its incidence is increasing in Korea [8]. The frequency of EGFR mutations in non-small cell lung cancer (NSCLC) has been reported to be 20-40% in Asian patients. We found that 24% of 104 randomly selected patients with lung adenocarcinoma carried mutations in the TK domain of the EGFR gene, which is consistent with previous reports [9-11]. Therefore, studies have confirmed that the EGFR mutation is common in Asian patients. Pao et al. [9] retrospectively examined 192 EGFR TK-domain mutations and found that 85.9% of these were in one of two hot spots. Similarly, 92% of the mutations in our study occurred in one of two hot spots.
It has been previously reported that EGFR mutations are more frequent in female patients and in those who have never smoked [7,12]. Shigematsu et al. [11] found EGFR gene mutations to be more prevalent in females than in males but independent of smoking status in Japanese patients. Huang et al. [13] reported no significant association between the presence of an EGFR mutation and gender or smoking history in Chinese patients. In the present study on Korean patients, EGFR mutations were not associated with gender, smoking history, age, histological grade, bronchioloalveolar components, or tumor stage. Although Korea is geographically close to China and Japan, many cultural differences, including smoking rate, nutrition, housing, and genetic backgrounds, exist among the three populations and may account for the different findings in these studies. Furthermore, differences may exist among the studies related to selection bias due to heterogeneous pathological profiles of patients [10].
It has been reported that EFGR mutations are detected more frequently in younger patients and in those with more advanced stages of lung cancer [9]. However, Kosaka et al. [14] found no association between EGFR mutations, age, and lung cancer stage. We also found that EGFR mutations were not associated with disease stage, suggesting that EGFR mutations occur relatively early in the clinical course of adenocarcinoma and are associated with its pathogenesis rather than its progression. EGFR mutations may also be more common in tumors with features of bronchioalveolar carcinoma (BAC) histology [6,7,10]. However, based on the World Health Organization's classification criteria for lung tumors [15], we found no association between EGFR mutation status and BAC or the BAC subtype of adenocarcinoma.
The reason for the marked difference in the response to anti-EGFR agents between East Asian and American patients is not clear. The geographical differences between the populations may reflect differences in genetic susceptibility. The first intron of the EGFR gene contains a dinucleotide (CA) repeat [16], and the number of CA repeats, which ranges from 14 to 21, correlates with the transcriptional activity [17,18]. Given this association between EGFR protein expression and the length of the CA repeat polymorphism, genotypic background may contribute to the variability in EGFR expression and thus to the differential response of patients to anti-EGFR agents [16,17]. The polymorphism lengths in East Asians are considerably longer than those in Caucasians and African-Americans. Furthermore, the difference in the incidence of non-smoking patients with lung cancer may in part reflect these genetic differences. We found that 85% of female patients and 26% of male lung cancer patients had never smoked. These percentages are much lower in American lung cancer patients, with only 15% of female and 6% of male patients with lung cancer in the U.S. having never smoked, and there is a similar trend in East Asia [18,19].
EGFR and KRAS mutations are mutually exclusive, suggesting that they have functionally equivalent roles in lung tumorigenesis [11,14]. Fifteen to 30% of lung adenocarcinomas contain activating mutations in a KRAS family member. These mutations occur most frequently in codons 12 and 13 encoded in exon 2 [21,22] and may be associated with unfavorable outcomes [23]. In the present study, a KRAS mutation was detected in 9.6% of the samples, which was a lower occurrence than that noted in other reports [24]. However, the frequency of the KRAS mutation among East Asian patients ranges from 8% to 13% of lung adenocarcinomas [10,14]. The high incidence of EGFR mutations in East Asians with lung adenocarcinomas may explain the lower frequency of KRAS mutations in East Asians compared with that in Caucasian patients. KRAS mutations more frequently occur in individuals with a history of substantial cigarette use and are increasingly likely to be a function of higher tobacco exposure [23]. Histological grade and cell type may be prognostic factors. Multiple studies have reported improved survival in never-smokers with primary adenocarcinoma of the lung [25,26]. De Perrot et al. [27] found that women were more likely to have never smoked than men and that 54% of women had the NSCLC subtype of adenocarcinoma. Furthermore, compared with men, women had superior survival independent of age, presence of symptoms, smoking habits, histology, or disease stage. Therefore, lung cancer has a different intrinsic biological behavior and natural history in women compared with men. The 5-yr survival rate for women with lung cancer was reported to be 17.3% compared with 13.8% for men [27].
In conclusion, we found a high prevalence of EGFR mutations in Korean patients with adenocarcinoma of the lung. EGFR mutations were never present in tumors that had KRAS mutations, indicating the possible importance of a genotypeoriented approach for diagnosis and treatment of pulmonary adenocarcinoma. Our results did not show a correlation between the presence of an EGFR mutation and smoking history, gender, age, or disease stage.
References
1. Manning G, Whyte DB, Martinez R, Hunter T, Sudarsanam S. The protein kinase complement of the human genome. Science 2002. 298:1912ŌĆō1934PMID : 12471243.


2. Arteaga CL. Overview of epidermal growth factor receptor biology and its role as a therapeutic target in human neoplasia. Semin Oncol 2002. 29(5 Suppl 14):3ŌĆō9PMID : 12422308.

3. Mendelsohn J. Targeting the epidermal growth factor receptor for cancer therapy. J Clin Oncol 2002. 20(18 Suppl):1SŌĆō13SPMID : 12235219.


4. Fukuoka M, Yano S, Giaccone G, et al. Multi-institutional randomized phase II trial of gefitinib for previously treated patients with advanced non-small-cell lung cancer. J Clin Oncol 2003. 21:2237ŌĆō2246PMID : 12748244.


5. Cappuzzo F, Gregorc V, Rossi E, et al. Gefitinib in pretreated non-small-cell lung cancer (NSCLC): analysis of efficacy and correlation with HER2 and epidermal growth factor receptor expression in locally advanced or metastatic NSCLC. J Clin Oncol 2003. 21:2658ŌĆō2663PMID : 12860941.


6. Paez JG, Janne PA, Lee JC, et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 2004. 304:1497ŌĆō1500PMID : 15118125.


7. Lynch TJ, Bell DW, Sordella R, et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 2004. 350:2129ŌĆō2139PMID : 15118073.


8. Kim YS, Kwon YS, Oh IJ, et al. National survey of lung cancer in Korea, 2005. J Lung Cancer 2007. 6:67ŌĆō73.

9. Tomizawa Y, Iijima H, Suguna N, et al. Clinicopathologic significance of the mutations of the epidermal growth factor receptor gene in patients with non-small cell lung cancer. Clin Cancer Res 2005. 11:6816ŌĆō6822PMID : 16203769.


10. Soung YH, Lee JW, Kim SY, et al. Mutational analysis of EGFR and K-RAS genes in lung adenocarcinomas. Virchows Arch 2005. 446:483ŌĆō488PMID : 15815931.


11. Shigematsu H, Lin L, Takahashi T, et al. Clinical and biological features of epidermal growth factor receptor mutations in lung cancers. J Natl Cancer Inst 2005. 97:339ŌĆō346PMID : 15741570.


12. Pao W, Miller VA. Epidermal growth factor receptor mutations, small-molecule kinase inhibitors, and non-small-cell lung cancer: current knowledge and future directions. J Clin Oncol 2005. 23:2556ŌĆō2568PMID : 15767641.


13. Huang SF, Liu HP, Li LH, et al. High frequency of epidermal growth factor receptor mutations with complex patterns in non-small cell lung cancers related to gefitinib responsiveness in Taiwan. Clin Cancer Res 2004. 10:8195ŌĆō8203PMID : 15623594.


14. Kosaka T, Yatabe Y, Endoh H, Kuwano H, Takahashi T, Mitsudomi T. Mutations of the epidermal growth factor receptor gene in lung cancer: biological and clinical implications. Cancer Res 2004. 64:8919ŌĆō8923PMID : 15604253.


15. Travis WD, Colby TV, Corrin B. Histological typing of lung and pleural tumours 1999. 3rd ed. New York: Springer-Verlag.
16. Ekstrand AJ, Sugawa N, James CD, Collins VP. Amplified and rearranged epidermal growth factor receptor genes in human glioblastomas reveal deletions of sequences encoding portions of the N- and/or C-terminal tails. Proc Natl Acad Sci U S A 1992. 89:4309ŌĆō4313PMID : 1584765.



17. Wong AJ, Ruppert JM, Bigner SH, et al. Structural alterations of the epidermal growth factor receptor gene in human gliomas. Proc Natl Acad Sci U S A 1992. 89:2965ŌĆō2969PMID : 1557402.



18. Toyooka S, Pass HI, Shivapurkar N, et al. Aberrant methylation and simian virus 40 tag sequences in malignant mesothelioma. Cancer Res 2001. 61:5727ŌĆō5730PMID : 11479207.

19. Minami Y, Tateno H. Associations between cigarette smoking and the risk of four leading cancers in Miyagi Prefecture, Japan: a multi-site case-control study. Cancer Sci 2003. 94:540ŌĆō547PMID : 14529588.


20. Cheng YW, Chiou HL, Sheu GT, et al. The association of human papillomavirus 16/18 infection with lung cancer among nonsmoking Taiwanese women. Cancer Res 2001. 61:2799ŌĆō2803PMID : 11306446.

21. Rodenhuis S, Slebos RJ, Boot AJ, et al. Incidence and possible clinical significance of K-ras oncogene activation in adenocarcinoma of the human lung. Cancer Res 1988. 48:5738ŌĆō5741PMID : 3048648.

22. Suzuki Y, Orita M, Shiraishi M, Hayashi K, Sekiya T. Detection of ras gene mutations in human lung cancers by single-strand conformation polymorphism analysis of polymerase chain reaction products. Oncogene 1990. 5:1037ŌĆō1043PMID : 2197591.

23. Rodenhuis S, Slebos RJ. Clinical significance of ras oncogene activation in human lung cancer. Cancer Res 1992. 52(9 Suppl):2665SŌĆō2669SPMID : 1562997.

24. Sekido Y, Fong KM, Minna JD. Progress in understanding the molecular pathogenesis of human lung cancer. Biochim Biophys Acta 1998. 1378:F21ŌĆōF25PMID : 9739759.


25. Nordquist LT, Simon GR, Cantor A, Alberts WM, Bepler G. Improved survival in never-smokers vs current smokers with primary adenocarcinoma of the lung. Chest 2004. 126:347ŌĆō351PMID : 15302716.


Figure┬Ā1
Mutations of the EGF receptor in lung cancer tissues. Most of mutations were found at Exon 19 and Exon 21. Numbers in parentheses indicated the number of lung cancer patients with mutations.
Table┬Ā2
Relationship between EGFR mutations and clinical/pathological features in a subset of patients




 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Download Citation
Download Citation Print
Print