LncRNA as a diagnostic marker for rheumatoid arthritis: a systematic review and meta-analysis
Article information
Abstract
Background/Aims
Rheumatoid arthritis (RA) is categorized as an autoimmune disorder characterized by the absence of a definitive cure. In this investigation, we executed a systematic review and diagnostic meta-analysis to ascertain the diagnostic efficacy of long non-coding RNAs (lncRNAs) in the identification of RA.
Methods
Our survey involved relevant studies published in PubMed, Web of Science, Embase, Cochrane library search engines. Specificity and sensitivity were calculated, as well as positive likelihood ratios (PLRs), negative likelihood ratios (NLRs), and diagnostic odds ratios (DORs). The operating characteristics of the overall receiver were plotted, and the area under the curve (AUC) was evaluated.
Results
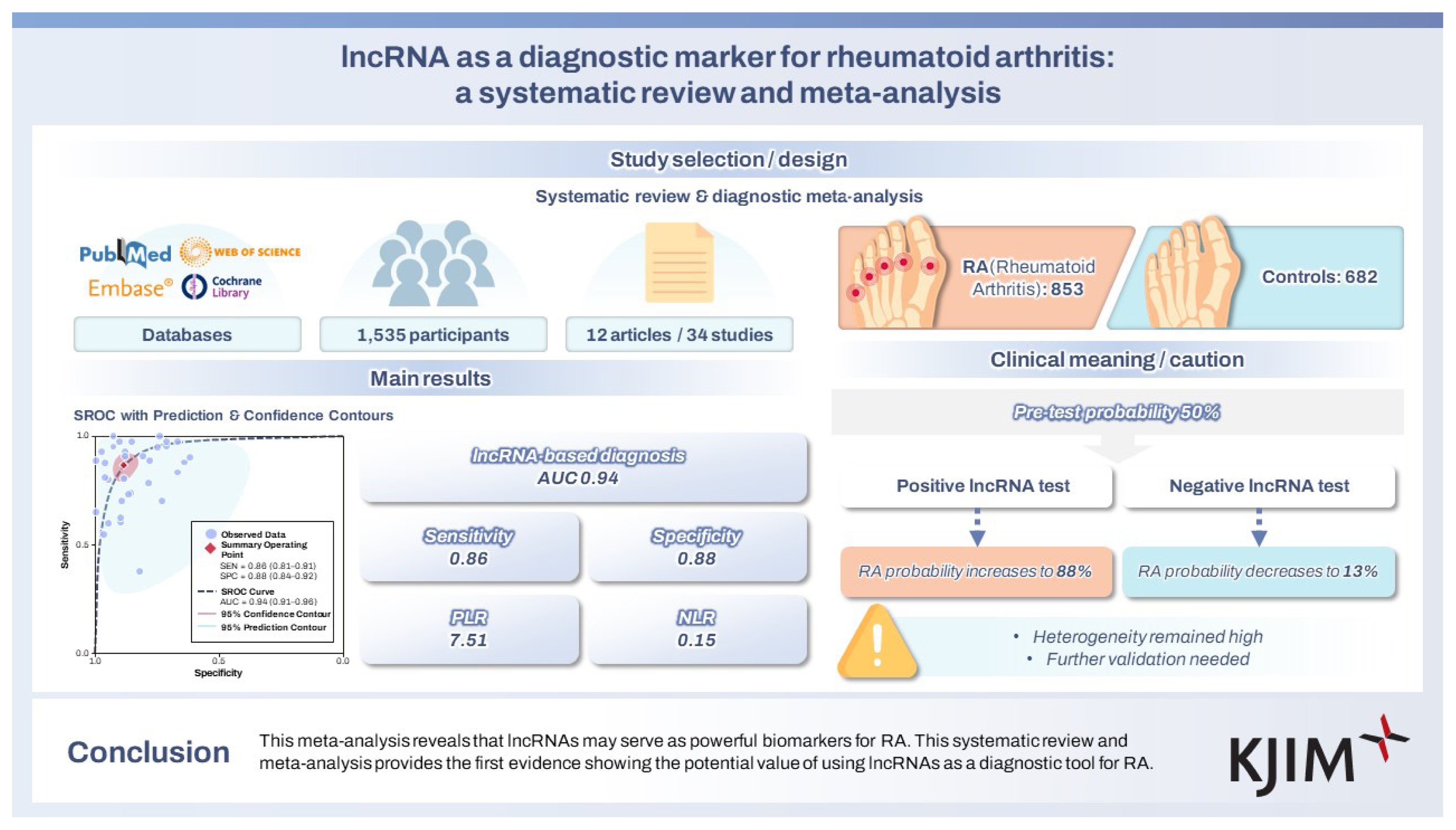
This systematic review and meta-analysis incorporated 34 studies involving a total of 1,535 participants—comprising 853 individuals diagnosed with RA and 682 control subjects without the disease. The pooled specificity, sensitivity, NLR, PLR, and DOR were 0.88 (95% CI: 0.84–0.92), 0.86 (95% CI: 0.81–0.91), 0.15 (95% CI: 0.11–0.22), 7.51 (95% CI: 5.39–10.46), and 49.06 (95% CI: 30.31–79.41), respectively, and the AUC = 0.94 (95% CI: 0.91–0.96). Subgroup analysis was performed according to lncRNA expression in RA, sample size, sample type, RNA extraction, and control group type.
Conclusions
This meta-analysis reveals that lncRNAs may serve as powerful biomarkers for RA. This systematic review and meta-analysis provides the first evidence showing the potential value of using lncRNAs as a diagnostic tool for RA.
INTRODUCTION
Rheumatoid arthritis (RA) is one of the autoimmune diseases affecting human health. One of its main pathological features is bone destruction, inflammation, and synovial hyperplasia [1], its global incidence is 0.21% to 0.51%. In recent years, relevant studies have found that the mortality rate of RA joints is decreasing, and the incidence increases with age [2,3]. The early diagnosis of RA mainly depends on the clinical manifestations of patients and the changes of sedimentation rate and complete blood count at the time of onset [4]. The most commonly used serum biomarkers include C-reactive protein, rheumatoid factor (RF), anti-cyclic citrullinated protein (anti-CCP), anti-cyclic citrullinated peptide 2 (anti-CCP2), anti-carp protein, 14-3-3η and other novel biomarkers, are also significantly manifested in RA patients [5–7]. The treatment and prognosis of RA still face challenges. Early diagnosis and intervention can effectively delay the progression of RA.
Long non-coding RNAs (lncRNAs) are a class of transcripts longer than 200 nt that lack protein-coding ability. LncRNAs play a pivotal regulatory role in the process of gene expression, cell biology, and carcinogenesis [8], rheumatoid diseases are similar to tumors, have a complex pathogenesis, unclear early diagnosis, and poor treatment outcomes, and drug treatment for rheumatoid diseases is initially also derived from cancer treatment [9]. LncRNAs are involved in regulating inflammatory responses and pathophysiological processes in RA, but are poorly conserved between species [10]. In the context of inflammatory diseases, lncRNAs are widely involved in various cellular processes such as apoptosis, metabolism, inflammation, cell differentiation, and proliferation [11]. LncRNAs are abnormally expressed in RA and are closely related to the occurrence and development of RA [12]. These studies lay the foundation for finding biological markers with strong specificity and high sensitivity. LncRNAs have promising applications as novel biological markers and therapeutic targets for RA. There have been several literature reports on lncRNAs as diagnostic markers RA, and the findings are inconsistent. Therefore, a thorough review of recommended biomarkers for RA diagnosis has long been warranted, and to our knowledge, this topic has not been systematically reviewed, and our goal is to perform a systematic review and meta-analysis of the diagnostic value of lncRNAs for the detection of RA in humans.
METHODS
In this study, we present a systematic review and meta-analysis of PICOS-based lncRNAs for their diagnostic value in RA. The PROSPERO registration number is CRD42024597196. Population (P): Participants in this systematic review and meta-analysis included individuals with a diagnosis of RA, while healthy people or those with isolated knee injuries who were clinically and serologically confirmed to have no history of RA or other systemic inflammatory diseases served as controls. Intervention (I): LncRNA detection and assessment study focused on evaluating lncRNA expression levels in RA patients and controls. According to Youden index, the test results were divided into positive and negative, and the critical threshold was established. Sensitivity and specificity were calculated using true positive (TP), true negative (TN), false positive (FP), and false negative (FN) values. Comparison (C): Non-RA controls compared the diagnostic performance of lncRNAs in RA patients with healthy individuals and untreated RA patients. American College of Rheumatology (ACR) diagnostic criteria served as the gold standard for validation. Results (O): The main objective of diagnostic performance was to determine the diagnostic value of lncRNAs in distinguishing RA patients from non-RA controls. Statistical analysis was performed to calculate pooled sensitivity, specificity, diagnostic odds ratio (DOR), and summary receiver operating characteristic (SROC) curve analysis to provide insights into diagnostic validity. Study design (S): This study followed the Preferred Reporting Items for Systematic Reviews and Meta-Analyses guidelines (PRISMA). Literature searches were conducted in the PubMed, Web of Science, Embase, and Cochrane Library databases in English language only. Search terms included “Arthritis, Rheumatoid”, “Noncoding RNA, Long”, “Diagnosis”, and “Biomarkers”. The most recent search update was on 22 April 2024.
Eligibility criteria
This study focused on the diagnostic value of lncRNA in RA patients and compared it with non-RA controls (healthy controls, treated RA patients, and isolated knee injuries without a history of RA and other systemic inflammatory diseases confirmed by clinical and serological tests). Inclusion criteria (1) The study was designed as a diagnostic test for lncRNA in the diagnosis of RA; (2) The data provided could deduce the sensitivity and specificity; (3) Each group contained more than 10 patients; (4) The study group was untreated RA patients; (5) The diagnostic criteria proposed by the ACR were used as the gold standard. Exclusion criteria: (1) short communication, case report, meeting, letter, book chapters, personal opinions, comments, and patents; (2) in vivo and in vitro studies; (3) duplicate studies; (4) studies with no full or incomplete information available or constructed 2 × 2 contingency tables; (5) studies written in languages other than English; (6) studies with insufficient data to identify FN, TN, FP, and TP values.
Study selection and data extraction
Two independent reviewers conducted literature screening based on predefined inclusion and exclusion criteria. Discrepancies were resolved through discussion or consultation with a third reviewer. Extracted data included author names, publication dates, and study characteristics.
Quality assessment
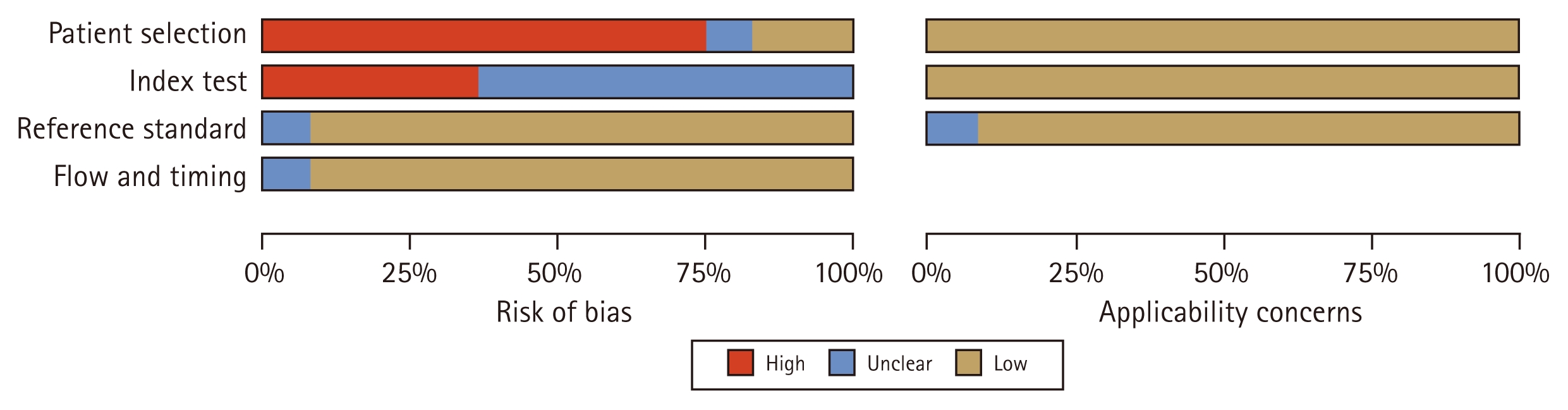
Study quality was evaluated using the Quality Assessment of Diagnostic Accuracy Studies-2 (QUADAS-2) tool. Two independent reviewers assigned responses as “Yes,” “No,” or “Unclear” for each quality criterion. Disagreements were resolved through consultation with a third reviewer, and assessments were performed using RevMan 5.4.1 software
Statistical analysis
Review Manager 5.4, Meta-Disc 1.4, Adobe Photoshop cc2018 and Stata17.0 software were used for statistical analysis. Combined sensitivity, specificity, positive likelihood ratios (PLR), negative likelihood ratios (NLR), and DOR with their corresponding 95% confidence intervals (CIs) were calculated, and comprehensive receiver operating characteristic curves (SROC) were plotted to calculate the area under the curve.
Heterogeneity analysis: threshold effect and non-threshold effect. Threshold effect: Heterogeneity caused by the threshold effect was considered when p < 0.05 by calculating the Spearman correlation coefficient between the logarithm of sensitivity and the logarithm of (1-specificity) and their p values. Non-threshold effect: Cochran’s Q test and I2 value were used to test heterogeneity caused by non-threshold effect. If I2 < 50%, it was considered that heterogeneity was small and fixed effect model was used; if I2 ≥ 50%, it was considered that heterogeneity was large and random effect model was used. In order to find the possible cause of heterogeneity, subgroup analysis and regression analysis were performed. Acceptable diagnostic yield by Fagan’s nomogram and publication bias by Deek’s funnel plot were plotted.
RESULTS
Basic information included in the study
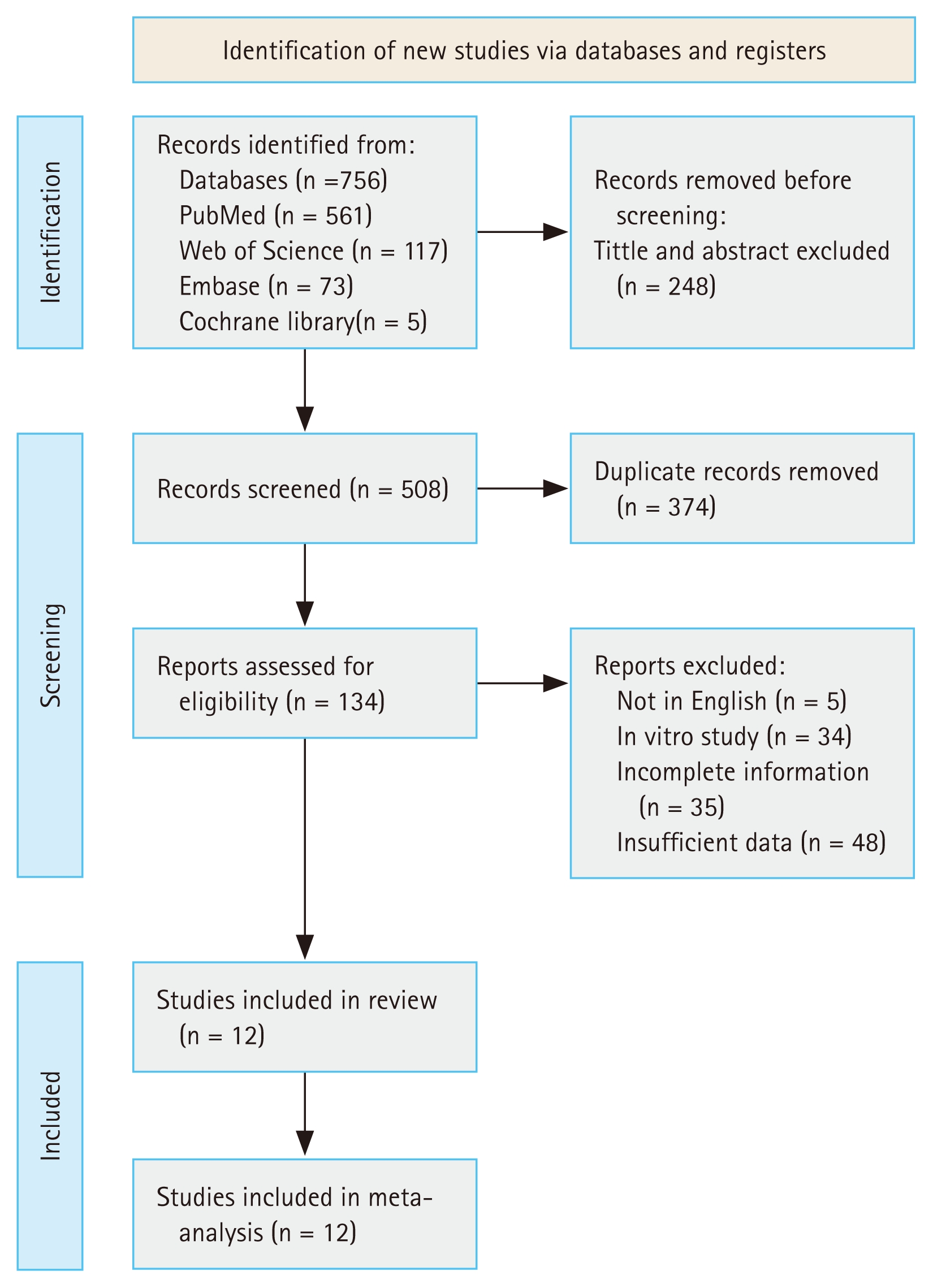
The search was performed in PubMed, Web of Science, Embase, and Cochrane library databases limited to English language with the search terms “Arthritis, Rheumatoid”, “Noncoding RNA, Long”, “Diagnosis”, and “Biomarkers”, and the last search was performed on April 22, 2024. A total of 756 relevant literatures were obtained, 248 repeated published literatures were excluded, and the remaining literatures were initially screened by reading the title and abstract according to the inclusion and exclusion criteria, and then further screened by carefully reading the full text to eliminate the literatures that did not meet the requirements, and finally 12 literatures [13–24] were included. The literature screening process and results of this study are shown in Figure 1.
Research retrieval and characteristics evaluation
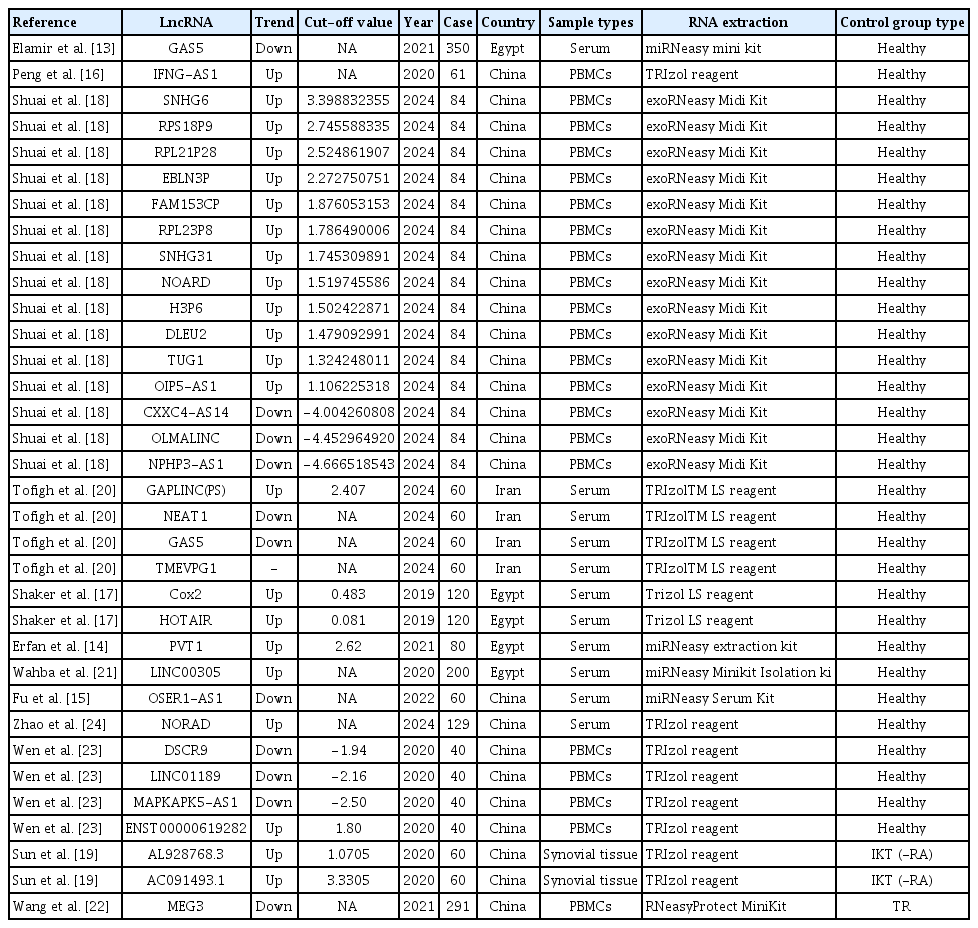
A total of 34 eligible studies were included in this meta-analysis to assess the screening and diagnostic value of lncRNAs for RA. The main characteristics of the included studies are listed in Table 1, and specific data details for subsequent analyses are presented in Table 2.
Research characteristics and quality evaluation
Twelve articles were finally included in this meta-analysis, and a total of 34 data were obtained to include the basic information of the studies (Table 1). QUADAS-2 was used for literature quality assessment. In this study, there was selection bias of cases, which was considered to be affected by the different sources of cases included in the articles, the majority of retrospective studies, and insufficient control of confounding factors. The risk of bias in terms of trials to be evaluated, gold standard, case flow and progression was low and clinical applicability was high (Fig. 2).
Heterogeneity test
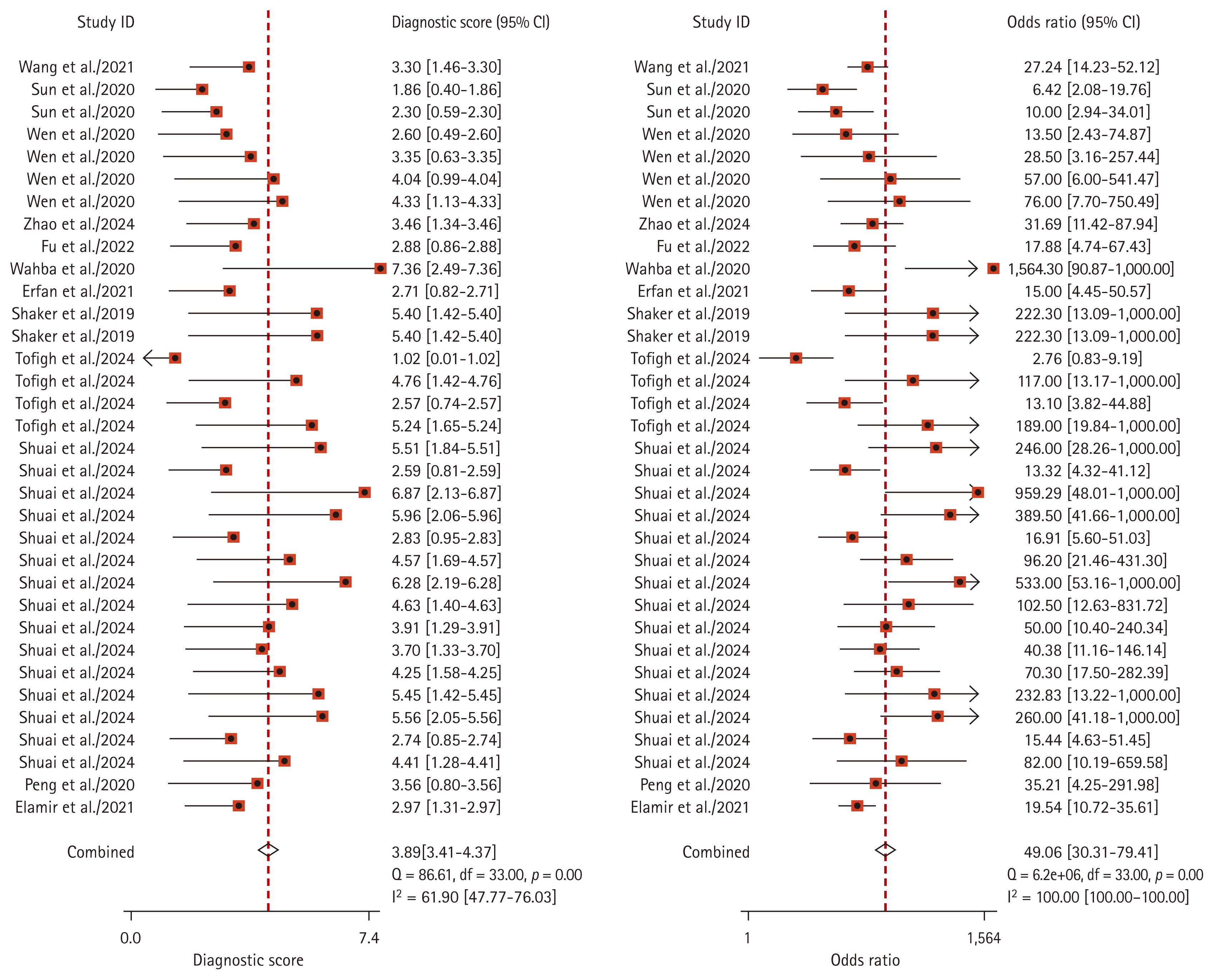
The included studies were tested for threshold heterogeneity by Meta-Disc 1.4 software with Spearman correlation coefficient p = 0.056, which showed p > 0.05 and no heterogeneity caused by threshold effects. Pooled sensitivity I2 = 85.30, p = 0.00; pooled specificity I2 = 78.83, p = 0.00; pooled PLR I2 = 65.39, p = 0.00; pooled NLR I2 = 84.91, p = 0.00; pooled DOR I2 = 100.00, p = 0.00; and pooled effect size I2 greater than 50%, p values less than 0.1 were calculated using the midas command in Stata17.0 software (Fig. 3–5). showed that there was heterogeneity between studies, and further analysis of the source of heterogeneity was required.
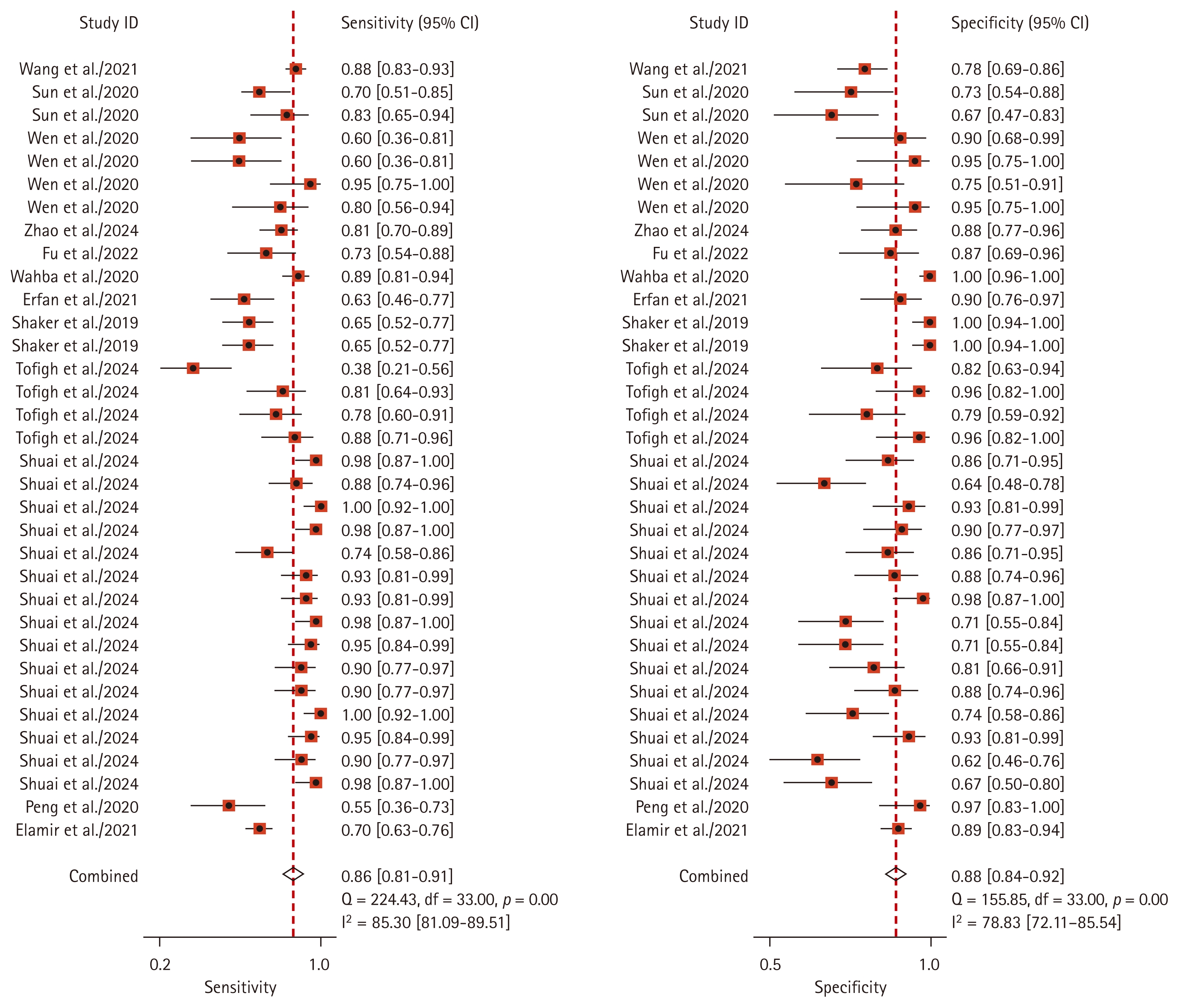
Sensitivity and specificity of lncRNA in the diagnosis of RA. LncRNA, long non-coding RNA; RA, rheumatoid arthritis; CI, confidence interval.
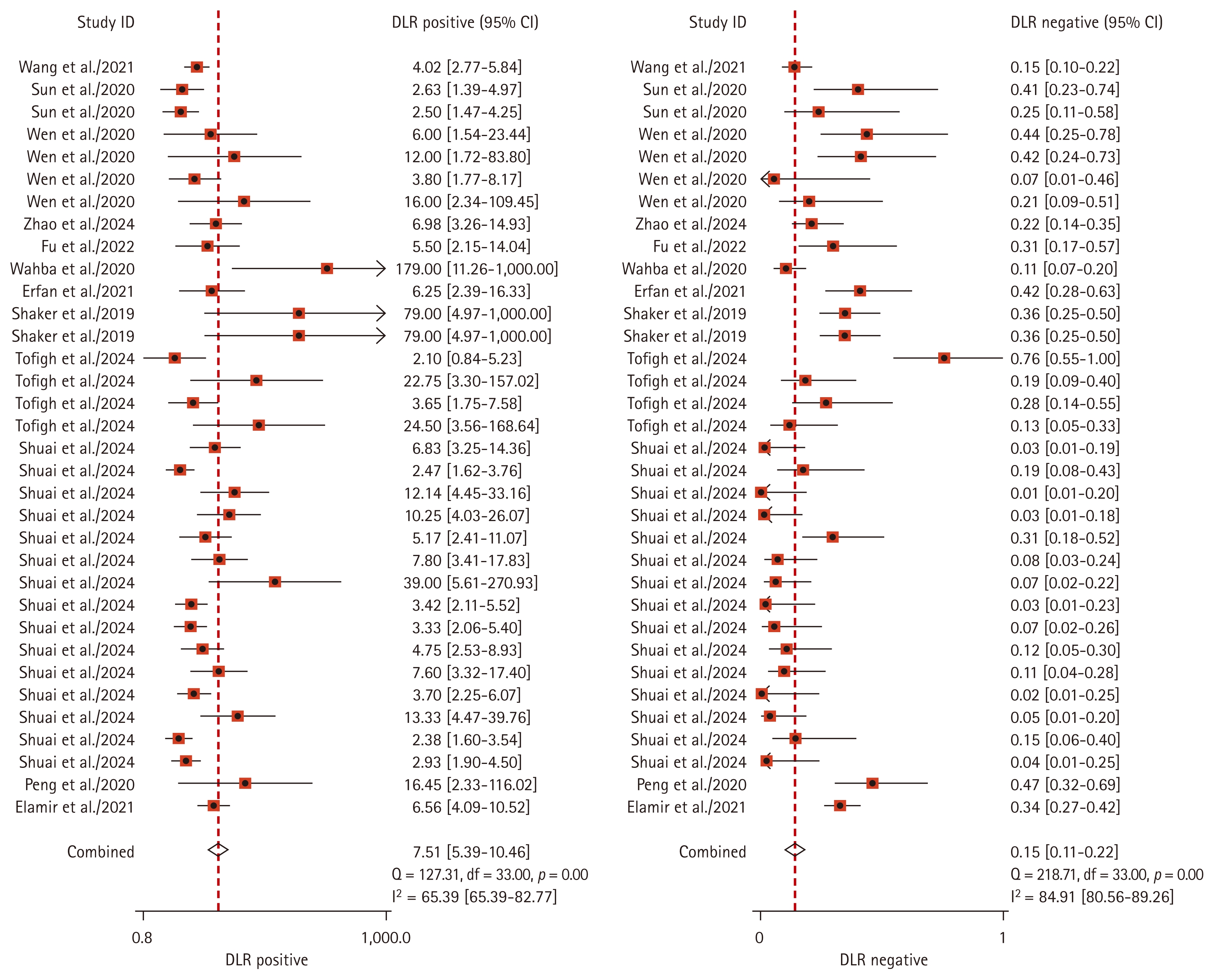
Forest plot of the combined likelihood ratio of lncRNAs diagnosis RA. LncRNA, long non-coding RNA; RA, rheumatoid arthritis; DLR, diagnostic likelihood ratio; CI, confidence interval.
Summary of diagnostic value of lncRNA biomarker
The pooled sensitivity of lncRNAs in the diagnosis of RA was 0.86 (95% CI: 0.81–0.91); specificity was 0.88 (95% CI: 0.84–0.92); PLR was 7.51 (95% CI: 5.39–10.46); NLR was 0.15 (95% CI: 0.11–0.22); and DOR was 49.06 (95% CI: 30.31–79.41). To assess diagnostic accuracy, a SROC curve was plotted with an AUC = 0.94 (95% CI: 0.91–0.96) (Fig. 3–6). The sources of heterogeneity were further explored by subgroup analysis and meta-regression analysis (Fig. 7, Table 2). Deeks’ funnel plot was used for publication bias analysis, and if the funnel plot was symmetrical, p > 0.05, it indicated that there was no publication bias (Fig. 8). Clinical value analysis was performed by plotting a Fagan plot, setting the pre-test probability and calculating the post-test probability (Fig. 9).
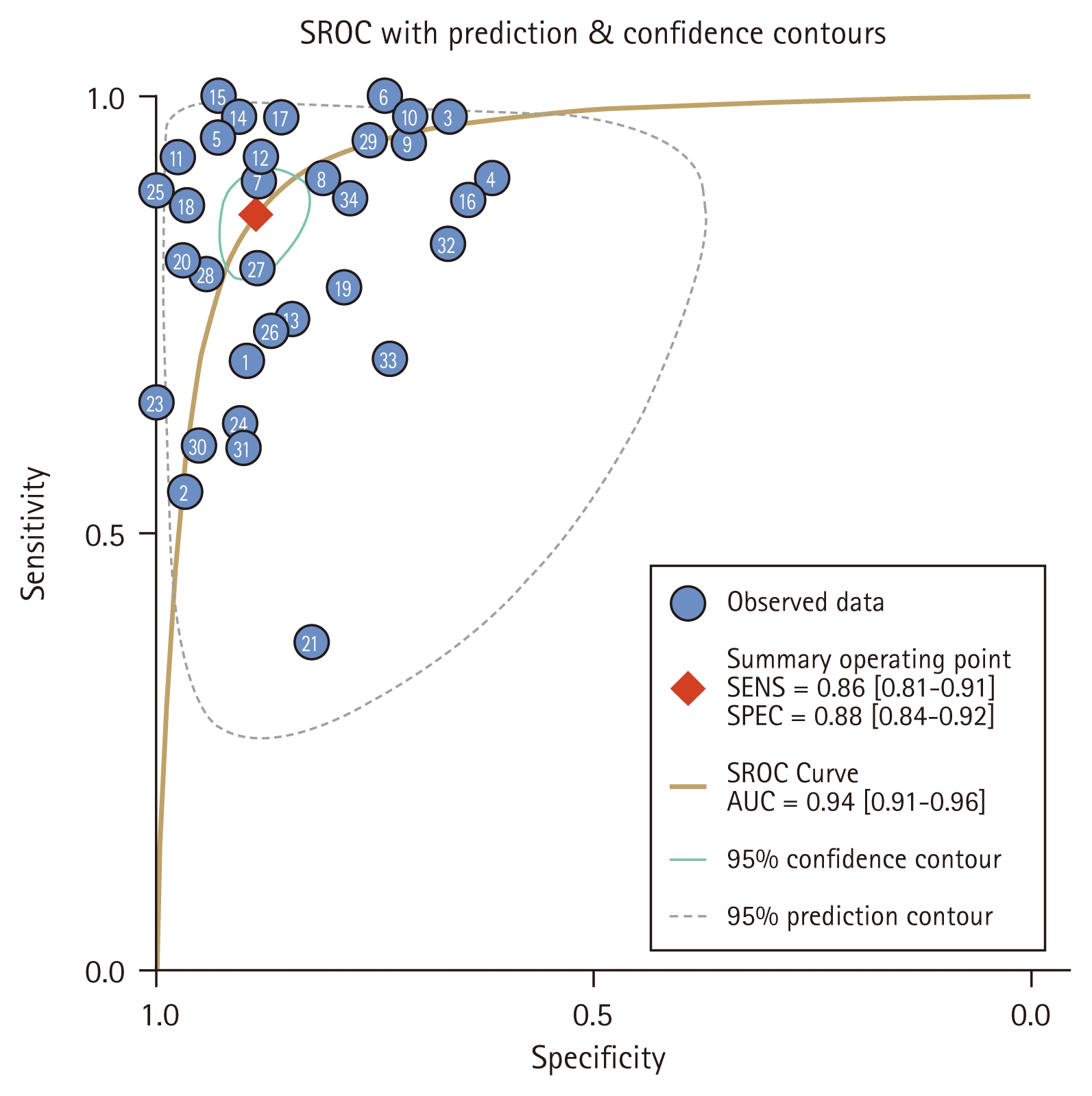
The SROC curves of lncRNAs diagnosis RA. SROC, summary receiver operating characteristic; LncRNA, long non-coding RNA; RA, rheumatoid arthritis; SENS, sensitivity; SPEC, specificity; AUC, area under the curve.
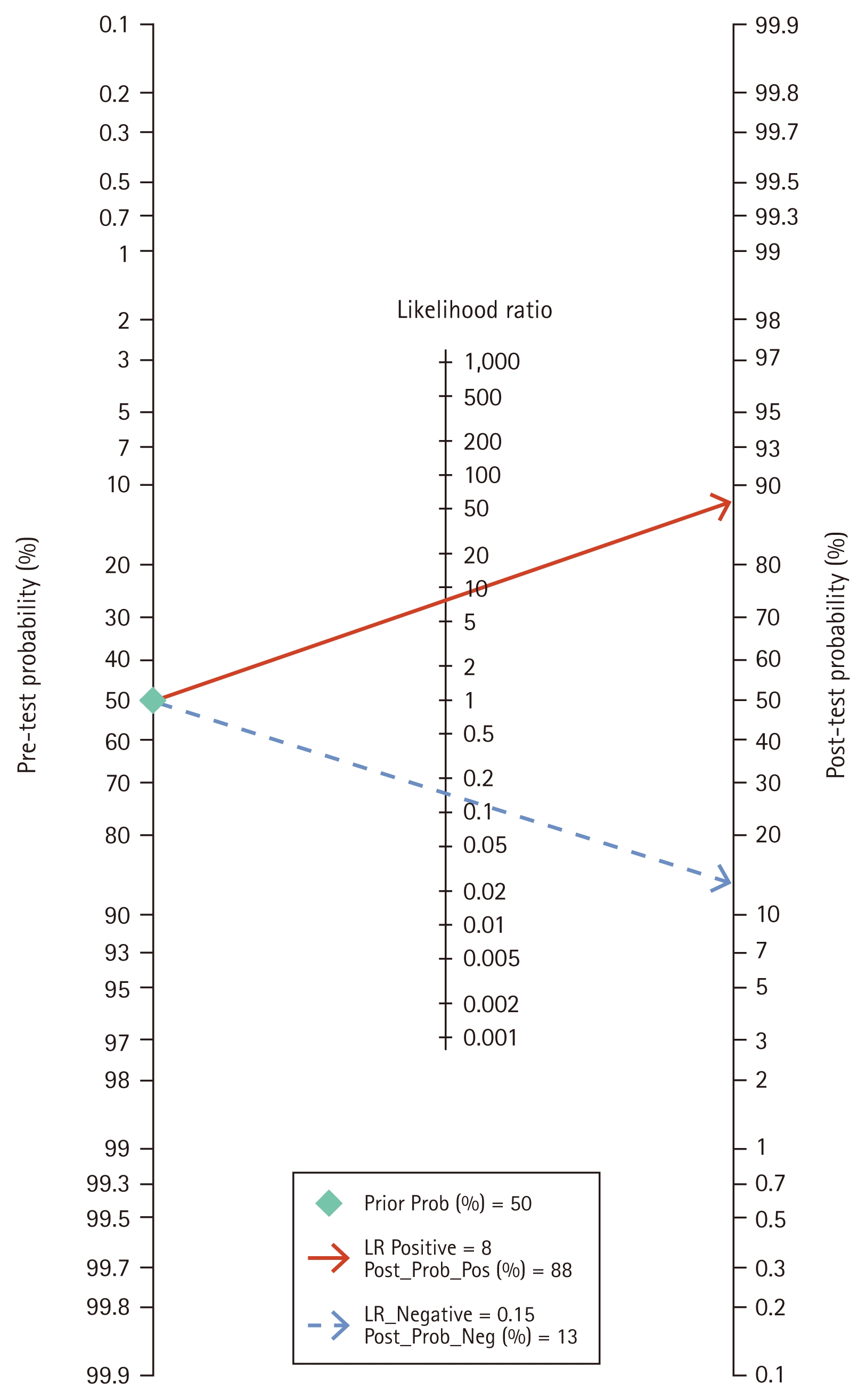
Fagan nomogram of lncRNAs diagnosis RA. LncRNA, long non-coding RNA; RA, rheumatoid arthritis; LR, likelihood ratio.
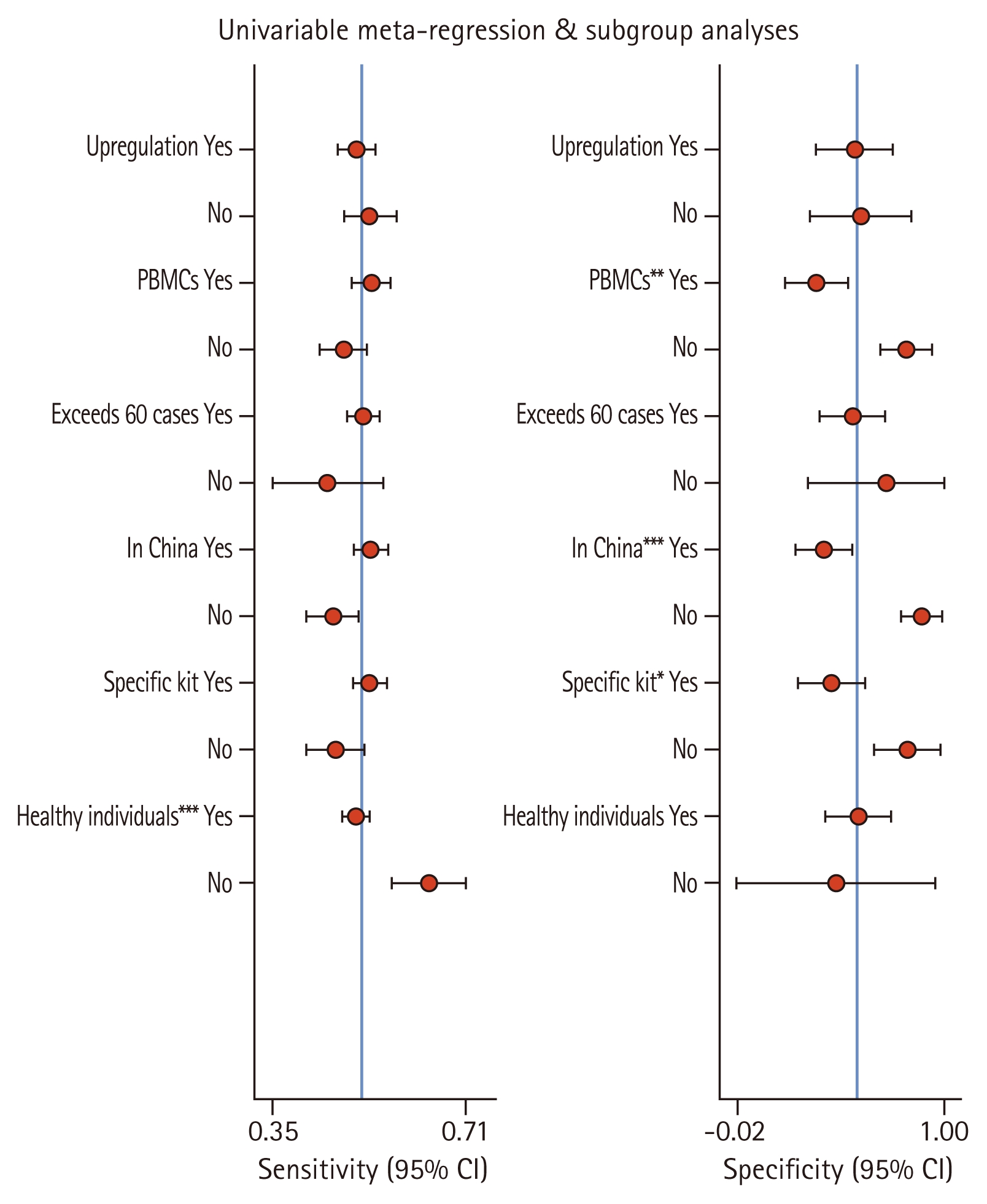
Meta regression and subgroup analysis
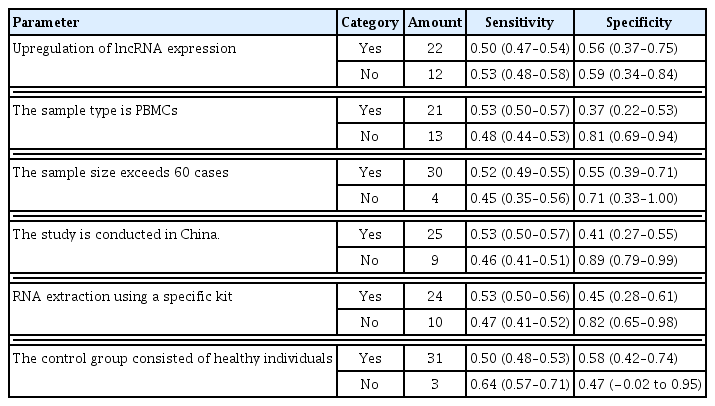
Because there was a large heterogeneity among the included studies, meta-analysis regression and subgroup analysis were used to further investigate the source of heterogeneity in this study. Subgroup analysis was performed according to whether lncRNA expression was increased in RA, whether the sample type was PBMCs, whether the sample size exceeded 60, whether the country where the experiment was performed was China, whether lncRNA was extracted with a specific kit, and whether the control group was a healthy population, and the results showed that the only statistically significant source of strong heterogeneity was “whether healthy individuals were used as controls” in terms of sensitivity (p < 0.001).In terms of specificity, the sources of heterogeneity were diverse, including “whether performed in China” (p < 0.001), “whether peripheral blood mononuclear cells (PBMCs) were used” (p < 0.01), and “whether specific kits were used” (p < 0.05).The sensitivity and specificity of decreased lncRNA expression in this included study were higher than that of increased lncRNA expression. The sample types used were mainly divided into PBMC, serum and synovial tissue. The results showed that the specificity of PBMC was higher than that of serum and synovial tissue, and the specificity of serum and synovial tissue was higher than that of PBMC. Sensitivity was higher for sample sizes of more than 60 and specificity was higher for sample sizes of no more than 60. Studies carried out in China were more sensitive than those in Iran and Egypt, and studies carried out in Iran and Egypt were more specific than those carried out in China. Specific kit extracted lncRNAs with higher sensitivity than Trizol reagent extracted lncRNAs, and Trizol reagent extracted lncRNAs with stronger specificity than specific kit extracted lncRNAs. Controls included healthy controls, treated RA patients, and independent knee traumas without RA, with treated RA patients and independent knee traumas without RA having higher sensitivity than healthy controls, and healthy controls having higher specificity than treated RA patients and independent knee traumas without RA (Table 2).
Clinical value analysis
Clinical value analysis was performed by plotting a Fagan plot, setting the pretest probability and calculating the posttest probability (Fig. 9), assuming a pretest probability of 50%, the probability of diagnosing RA increased from 50% to 88% when the long non-coding value was above the cutoff value, and decreased from 50% to 13% when the lncRNA value was below the cutoff value (Fig. 8). Thus, lncRNA can improve the diagnostic rate of RA and has certain clinical value.
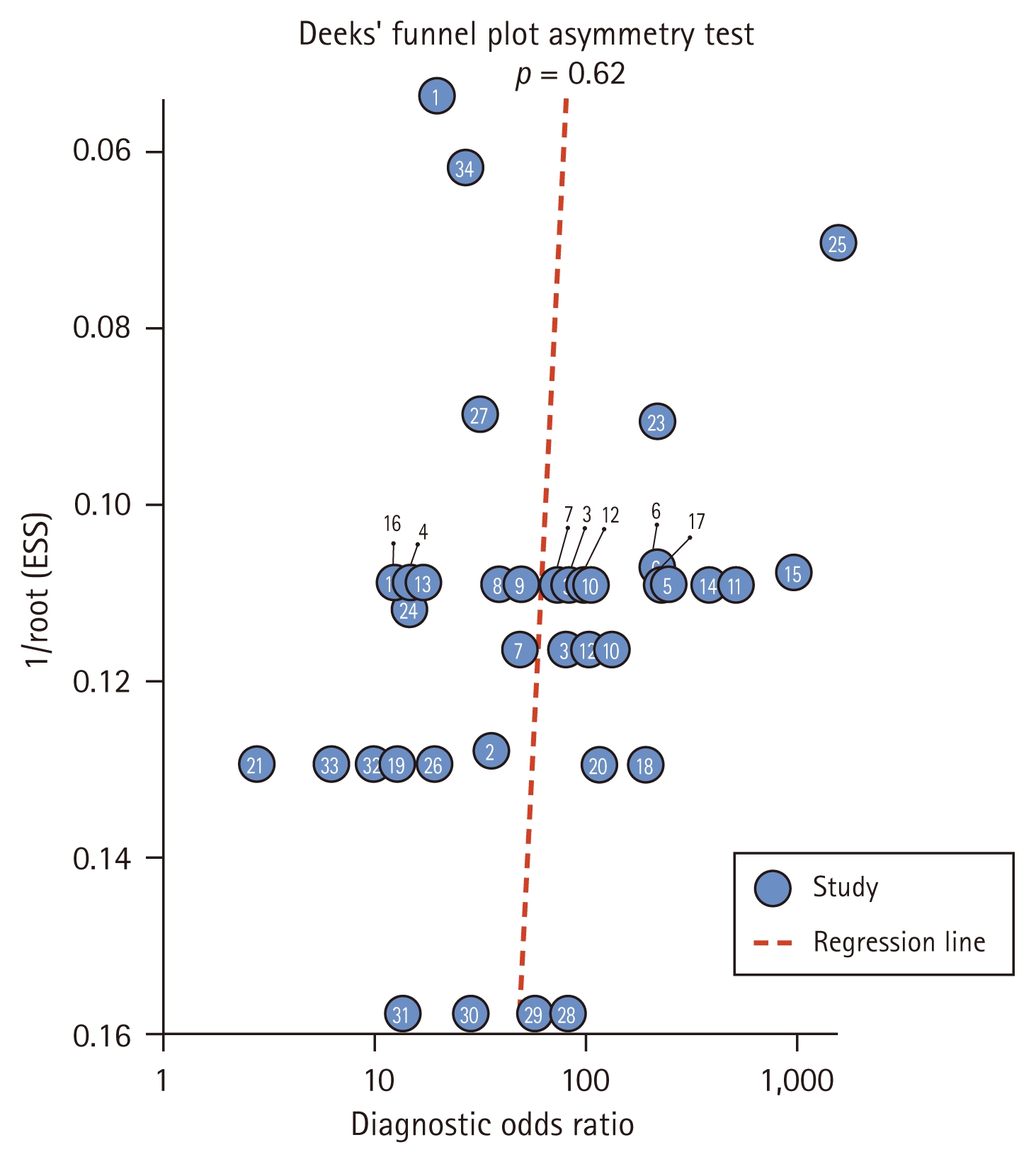
Publication bias
The Deeks’ funnel plot distribution plotted in this study was symmetrical with a slope coefficient of p = 0.62, indicating the absence of publication bias (Fig. 9).
DISCUSSION
The pooled sensitivity of lncRNA in the diagnosis of RA was 0.86 (95% CI: 0.81–0.91); specificity was 0.88 (95% CI: 0.84–0.92); PLR was 7.51 (95% CI: 5.39–10.46); NLR was 0.15 (95% CI: 0.11–0.22); and DOR was 49.06 (95% CI: 30.31–79.41); AUC = 0.94. In this study, we grouped as “Whether lncRNAs are up-regulated in RA”, “whether the sample type is PMBCs”, “whether the sample size exceeds 60”, “whether the country in which the experiment is carried out is China”, “whether lncRNAs are extracted with specific kits”, and “whether the control group is a healthy population” for meta-regression and subgroup analysis. The results of this meta-analysis did not prove that lncRNAs are good biological markers for the diagnosis of RA, but certain specific lncRNAs are potential indicators of clinicopathological features of RA. As a common serious immune system disease, RA seriously affects the daily work, daily activities and physical and mental health of patients [1,25]. Finding new biomarkers for RA is particularly important for the early diagnosis and prognostic evaluation of RA. LncRNAs are a class of transcripts greater than 200 nt in length that lack protein-coding capacity and play critical regulatory roles in RA. The pathological process of RA involves dysfunction of various immune cells such as T cells, B cells, macrophages and dendritic cells. LncRNAs play an important role in regulating immune cell function and differentiation [26]. Some studies have found that RNU12 affects T cell cycle involvement in RA by targeting C-JUN and CCNL2 [27]. NEAT1 is an accessory active molecule for CD4+ T cell differentiation into Th17 cells, and knockdown of NEAT1 actively inhibits Th17/CD4+ T cell differentiation by decreasing STAT3 protein levels [28]. Several specific lncRNAs chains exhibit strong regulatory functions in cell physiology and maturation of dendritic cells, such as lncTAF15:1-1 coordinating dendritic cell migration and CD4+ T cell recruitment by activating the PI3K/AKT/mTOR pathway to enhance CCL5 secretion [29]. LncRNAs such as Linc00324 and NEAT1 were found to be upregulated upon pro-inflammatory stimulation, involved in regulating signaling pathways such as NF-κB and MAPK, and enhancing the release of inflammatory factors such as TNF-α, IL-6, and IL-1β [25, 30]. LncRNA H19 is highly expressed in RA macrophages and promotes M1 macrophage activity and macrophage migration by up-regulating KDM6A [31]. Specific lncRNAs enhance the proliferative capacity and migratory activity of synovial fibroblasts through multiple molecular mechanisms in promoting their invasiveness. Elevated GAPLINC expression promotes tumor-like biological features of RA-FLS, and targeting GAPLINC significantly impairs the ability of RA-FLS cells to proliferate, migrate, and invade [32]. Lnc-AL928768.3 promotes proliferation, invasion ability as well as release of inflammatory factors in FLS from RA patients and inhibits apoptosis of RA-FLS by activating LTB-mediated NF-κB signaling pathway [33]. In addition, some researchers have found that lncRNAs strands are involved in inflammatory factor storms, and certain lncRNAs can regulate key inflammatory signaling pathways through sponge mechanisms during the formation of inflammatory factor storms. LncRNA H19 is highly expressed in MH7A cells, which can promote the secretion of IL-6 and TNF-α by activating NF-κB and JNK/p38 MAPK pathway signaling pathways, thereby participating in the formation of inflammatory factor storms and exacerbating joint lesions [34]. ZNF667-AS1 plays a protective role during RA development by sponging miR-523-3p and inactivating JAK/STAT signaling [35]. LncRNA MEG3 may regulate SMAD7 expression through a competitive sponge of miR-93-5p, thereby mediating inflammatory and proliferative phenotypes of RASFs, as well as enhanced infiltration of immune cells in the RA synovium [36]. In pathological conditions, dysregulation of lncRNAs is often accompanied by increased osteoclast numbers and enhanced bone resorption activity, leading to severe bone mass loss and structural destruction. Certain specific lncRNAs play key roles in the differentiation and function of osteoclasts, which in turn affect the homeostasis balance of bone tissue. It has been shown that lncRNA-Gm5532, which acts as a competitive endogenous RNA to regulate osteoclast differentiation [37]. KCNQ1OT1 regulates osteoclast differentiation through miR-128-3p/NFAT5 axis [38]. As one of the hotspots of molecular biology research in recent years, lncRNAs have shown important potential in the pathogenesis and clinical diagnosis of a variety of diseases. The results of this meta-analysis could not demonstrate that lncRNAs are good biomarkers for the diagnosis of RA, but evidence suggests that specific lncRNAs play a key regulatory role in pathological processes such as immune regulation, inflammatory response, and abnormal proliferation of synoviocytes, suggesting that they may become novel biomarkers in the diagnosis of RA. Currently, the classification criteria for RA jointly developed by the ACR and the European League Against Rheumatism (EULAR) mainly rely on clinical manifestations, imaging changes, and serological markers such as RF and anti-CCP [39]. However, these criteria have the problem of insufficient sensitivity in seronegative RA patients or early stages of the disease, making timely diagnosis difficult in some patients. In this context, the introduction of lncRNAs is expected to complement the limitations of existing diagnostic systems. Studies have shown that some lncRNAs present specific expression patterns in peripheral blood or synovial tissues of RA patients, and their expression levels are closely related to disease activity (DAS28 score), inflammatory parameters and the degree of joint destruction. Therefore, detecting the expression levels of these lncRNAs could provide molecular level support for the auxiliary diagnosis of RA, especially in patients who are negative for both RF and anti-CCP, lncRNAs may fill the gap in serological testing [40]. In addition, lncRNAs have also shown potential value in the recognition of early RA. Because early symptoms of RA are often atypical and easily confused with other arthritides, traditional serological markers tend to be positive after the disease progresses to a certain stage, resulting in delayed diagnosis. However, some lncRNAs showed significant up-regulation or down-regulation at the early stage of the disease, with high tissue specificity and stability, and could be detected non-invasively by blood samples, so as to achieve earlier intervention and treatment. In RA patients, alterations in lncRNA expression provide an opportunity to develop new therapeutic targets or biomarkers [26]. Although lncRNAs still face challenges such as standardization of detection techniques, cost control, and large-scale validation in clinical applications, they are gradually moving from basic research to translational medicine as potential tools to supplement existing ACR/EULAR criteria and open up new paths for the accurate diagnosis of RA.
Limitations
The disadvantage of this study is that: (1) studies published on other platforms or in other languages that meet the inclusion criteria may be missed; (2) Heterogeneity exists among studies, which may be caused by different lncRNA types. Due to the limited number of studies, detailed subgroup analysis is not possible; Due to the limited number of literatures, this study could not identify a specific lncRNA type for the diagnosis of RA. (3) All the original data included in this meta-analysis were derived from specific populations in East Asia (China), Middle East (Iran), and North Africa (Egypt). Although this provides ethnic diversity to some extent, this diversity is limited. Within these three countries, there is also diversity of ethnic subtypes, and available data may not adequately capture differences at this level. There may be differences in genetic background, lncRNA expression profiles, and disease heterogeneity among ethnic groups, which may affect the generalizability and performance of our diagnostic model in other ethnic groups worldwide. In summary, lncRNA as a novel biomarker of RA has gradually shown its potential to outperform traditional markers of RA. However, there are still huge challenges before lncRNAs can be used in clinical diagnosis. Although many literatures have reported that lncRNA is associated with various stages of the occurrence and development of RA, the underlying biological mechanism needs to be further clarified. Exploring the mechanism of action of lncRNA in RA, unifying technical standards for detection or constructing a new diagnostic model combined with other biological markers may enhance the diagnostic value of lncRNA as a biomarker for RA.
Conclusion
This meta-analysis reveals that lncRNAs may serve as powerful biomarkers for RA. This systematic review and meta-analysis provide the first evidence showing the potential value of using lncRNAs as a diagnostic tool for RA. At present, numerous related large-scale studies conducted in different countries have continuously verified the efficacy of lncRNAs in the diagnosis of RA. However, there are still significant challenges in their clinical application. RA is a disease with multi-stage and heterogeneous characteristics, and existing studies often lack tracking analysis of the dynamic changes of lncRNAs with disease progression and treatment response. Combined with the different stages of the disease (early and late) and treatment response, the dynamic expression pattern of lncRNA can be traced for a long time to identify its diagnostic value in different stages of RA. In addition, lncRNA is expected to become an important supplement for accurate diagnosis of RA by combining with immune markers and other molecular indicators. Targeting the functional regulation of key lncRNAs in RA pathology may also open up new therapeutic avenues.
KEY MESSAGE
1. Integrate multiple sets of raw data through meta-analysis to quantitatively validate the overall diagnostic efficacy of lncRNAs for RA.
2. Effectively compensate for the diagnostic limitations of traditional markers such as RF and anti-CCP.
3. Providing a basis for standardizing clinical lncRNA-based diagnosis of RA in the future.
Notes
CRedit authorship contributions
Lin-ling Luo: formal analysis, writing - original draft; Yu-tong Chen: resources, software; Yi-ning Xu: data curation, supervision; Miao Lu: writing - original draft, visualization; Xi-de Liu: writing - review & editing, funding acquisition
Conflicts of interest
The authors disclose no conflicts.
Funding
This research was supported by Zhejiang Provincial Natural Science Foundation of China under Grant (No. LY21H270001) and Joint TCM Science & Technology Projects of National Demonstration Zones for Comprehensive TCM Reform (No. GZY- KJS-ZJ-2025-085) and Hangzhou Medical and Health Science and Technology Project (A20250378).