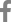|
 |
| Korean J Intern Med > Volume 22(4); 2007 > Article |
|
Abstract
Background
The expression of c-FLIP (cellular Fas-associated death domain-like interleukin-1 ╬▓-converting enzyme (FLICE)-inhibitory protein), which is a member of the family of inhibitors of apoptosis, has been associated with tumor development and progression. The aim of this study was to evaluate the expression of c-FLIP in gastric cancer and its correlation with tumor cell proliferation, apoptosis and the clinicopathologic features.
Methods
Immunohistochemical staining with anti-c-FLIP antibody was performed in 98 tissue samples obtained from gastric cancer patients who underwent surgical treatment. The apoptotic cells were visualized by terminal deoxynucleotidyl transferase (TdT) mediated deoxyuridine triphosphate (dUTP) nick-end labeling (TUNEL), and the proliferative cells were visualized by staining with Ki-67 antibody.
Results
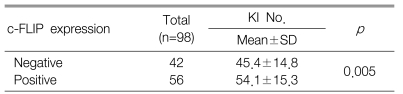
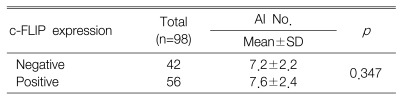
The positive expression of c-FLIP in the gastric cancer tissues was demonstrated in 57.1% of the cases. The expression of c-FLIP was increased in the gastric cancer tissues compared with the matched normal gastric mucosa. The expression of c-FLIP was significantly associated with histologic differentiation (p=0.038). However, there was no association between the c-FLIP expression and the other clinicopathological parameters, including patient survival. The Ki-67 labeling index (KI) for the 98 tumors ranged from 7.6 to 85.0 with a mean KI of 50.4┬▒15.7. The mean KI value of the c-FLIP positive tumors was 54.1┬▒15.3 and this was significantly higher than that of the c-FLIP negative tumors (p=0.005). The apoptotic index (AI) for the 98 tumors ranged from 0.0 to 10.0 with a mean AI of 7.4┬▒2.3. There was no significant difference between the c-FLIP expression and the AI (p=0.347).
Apoptosis plays an important role in organ homeostasis by eliminating senescent or damaged cells. Disturbance of apoptosis may confer a survival advantage on malignant cells that harbor genetic alterations and thus promote neoplastic progression1-3). Apoptosis induction may occur by two distinct signaling pathways. In the extrinsic pathway, apoptosis is induced through the binding of a ligand to a death receptor on the cell surface, whereas in the intrinsic pathway, apoptosis is mediated by mitochondrial release of cytochrome c. The central event in apoptosis is the proteolytic activation of a class of cysteine aspartyl-specific proteases termed the caspases. Initiator caspases cleave effector caspases, which in turn degrade a number of intracellular protein substrates and thereby induce the characteristic morphological hallmarks of apoptosis. Specialized initiator caspases are located at the top of both apoptosis signaling cascades. Caspases-8 and -10 are mediators of the extrinsic pathway, and caspase-9 initiates the intrinsic pathway3, 4).
Cellular Fas-associated death domain-like interleukin-1 ╬▓-converting enzyme (FLICE) inhibitory protein (c-FLIP) is a catalytically inactive caspase-8 and caspase-10 homologue, and its gene was recently identified to be a novel antiapoptotic gene5-8). c-FLIP is recruited to death receptor signaling complexes, where it inhibits the caspase activation that's responsible for apoptosis and this activation is induced by diverse triggering stimuli5-8). The expression of c-FLIP in the vast majority of human cancers, including gastric cancer, is enhanced and associated with tumor development and progression9-13).
Regarding tumor cell proliferation, it is widely accepted that the proliferative capacity may influence the clinical course and hence, the patient prognosis. Ki-67 is a nuclear antigen that's expressed in all stages of the cell cycle, except G0 and early G1. In numerous studies, Ki-67 is an established proliferation marker and it has been extensively used to estimate the growth fraction of tumors14, 15).
Tumor development and progression results from the imbalance between cell proliferation and cell death, and most of this occurs through apoptosis16). The c-FLIP expression is believed to be involved in tumor development and progression by affecting cell proliferation and apoptosis. So, the aim of this study was to evaluate the expression of c-FLIP in gastric cancer and to determine its correlation with tumor cell proliferation, apoptosis and the clinicopathologic features.
We studied the formalin-fixed and paraffin-embedded tumor specimens from 98 randomly chosen patients who had undergone surgery for gastric cancer at Chonnam National University Hospital between July 1993 and June 1994. None of the patients had received preoperative radiotherapy or chemotherapy. The pathologic reports and clinical histories at the time of surgery were reviewed from the medical records. The histologic grade was classified according to the criteria of Lauren and the World Health Organization17, 18). The tumor staging was in accordance with the American Joint Committee on Cancer (AJCC) staging system19). Survival was measured from the time of surgery until follow-up at September 2003. This study group was comprised of 66 males and 32 females. The median age was 60.0┬▒10.2 (mean┬▒SD) with an age range from 33 to 79 years. The mean size of tumor was 5.1┬▒2.5 cm (mean┬▒SD) with a range from 0.5 to 13.0 cm. The mean follow-up period was 75.9 months with a range from 1.3 to 119.8 months
All procedures for the immunohistochemical staining were done by the Micro-Probe staining system, which is based on capillary action20). Paraffin sections 4 ┬Ąm in thickness with a mounted probe on the slides were immunostained with anti-rabbit polyclonal antibody for the c-FLIP antigen (R&D systems, Inc., NJ, USA) by the avidin-biotin peroxidase complex method. The sections were deparaffinized and heated in a microwave oven for 7 minutes to retrieve the antigens. They were immersed in 0.6% hydrogen peroxide for 10 minutes to block the endogenous peroxidase activity. The primary antibody, at concentration of 1:100, was diluted in phosphate-buffered saline supplemented with 5% normal horse serum and 1% bovine serum albumin, and then this was incubated with the tissues for overnight at room temperature. Anti-mouse immunoglobulin G (Sigma, St. Louis, MO) labeled with biotin was used as a secondary antibody for the detection of the primary antibodies, and the slides were incubated for 10 minutes at 45Ōäā with this secondary antibody. After multiple rinses with universal buffer, the streptavidin-alkaline phosphatase detection system (Biomeda, Foster, CA) was applied for 8 minutes. As the final step, the slides were developed for 10 minutes with the enzyme substrate 3, 3-diaminobenzidine (DAB, Sigma, St. Louis, MO). The slides were counterstained with hematoxylin solution for 3 minute (Research Genetics, Huntsville, AL). After dehydration, the tissue was sealed with a universal mount (Research Genetics, Huntsville, AL). For negative controls, the primary antibody was omitted and replaced with phosphate-buffered saline.
To quantiate the FLIP expression, a scoring method was modified from that by Zhou et al13). The staining intensity was classified from zero (no staining) to 3 (strong staining), and the percentage of the staining area was classified as 0 for no positive staining of tumor cells, 1 for positive staining in <10% of the tumor cells, 2 for positive staining in 10% to 50% of the tumor cells, or 3 for positive staining in >50% of the tumor cells. A staining index was calculated as the product of the staining intensity and the staining area. Assessment of the staining was evaluated by two independent pathologists who were without knowledge of the clinical outcomes such as the tumor stage, grade and survival. Consensus scores were assigned for each case by reviewing the slides that showed discrepancies in scoring. All sections for which the two observers disagreed were re-evaluated and discussed. There was total agreement on the classification. The tumors were categorized as having a positive expression (staining index Ōēź4) or a negative expression (staining index <4).
The tumor proliferative cells were visualized by immunohistochemically staining them using anti-Ki-67 antibody (MIB-1; diluted 1:150; Dakopatts, Glostrup, Denmark). Distinct nuclear immunoreactivity for Ki-67 was considered positive. The Ki-67 labeling index (KI) was presented as the number of Ki-67-positive nuclei per 1000 tumor cell nuclei. The determination of KI has been used to estimate the proliferative ability of tumor cells.
Apoptotic cells were detected by terminal deoxynucleotidyl transferase (TdT)-mediated deoxyuridine triphosphate (dUTP) nick end labeling (TUNEL)21). The TUNEL method for the detection of apoptotic cells in the 4 ┬Ąm thick sections of the formalin-fixed and paraffin-embedded tissue was performed by using the ApopTagŌäó Plus In Situ Apoptosis Detection Kit (Intergen, Purchase, NY). Briefly, the tissue sections were dewaxed and hydrated in a graded series of alcohol solutions. After they had been digested with proteinase K (20 mg/mL; Sigma, St. Louis, MO) for 15 min at room temperature, the endogenous` peroxidase was blocked by 2% hydrogen peroxide (H2O2) in phosphate buffered saline. TdT incubation was then performed for 120 min at 37Ōäā. The reaction was terminated by prewarmed working strength STOP/WASH buffer and then the sections were incubated with anti-digoxygenin peroxidase for 60 min at room temperature. After the DAB color reaction, nuclear counterstaining was performed with Mayer's hematoxylin. The positive control sections were treated with 0.7 g/mL DNase I (Sigma, St. Louis, MO) in potassium cacodylate buffer (pH 7.2) for 10 min before treatment with TdT buffer. The negative control sections were treated by substituting distilled water for working strength TdT enzymes. A quantitative method for calculating apoptotic cells was used by two independent pathologists. All the sections were examined in high power fields (magnification of 40├Ś10). The fields were selected in the highest labeled area per case. The apoptotic indices (AI) were expressed as a number of positive nuclei, including apoptotic bodies, among 1000 tumor cell nuclei.
All statistical analyses were carried out using the Statistical Package for the Social Sciences (SPSS/PC+ 13.0, Chicago, IL). The correlation between the c-FLIP expression and the clinicopathological parameters was examined by Žć2-testing and Fisher's exact test. The relationship between the c-FLIP expression and the KI or AI was evaluated by the student t test. Survival curves were calculated according to the Kaplan-Meier method and the differences were tested with a log-rank test. The Cox regression model was used to determine the prognostic significance of each parameter by performing multivariate analysis. A p-value of less than 0.05 was considered statistically significant.
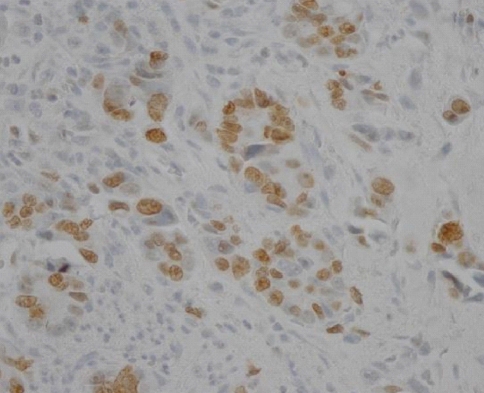
The immunostaining of c-FLIP was predominantly identified in the cytoplasmic regions of cancer cells and it was also detectable in the normal gastric mucosa, while it was not detectable in the stromal compartment (Figure 1A, B). The expression of c-FLIP was increased in the gastric cancer tissues compared with the matched normal gastric mucosa. Based on our criteria, the positive expression of c-FLIP in the cancerous tissues was 57.1% (56/98). Ki-67 immunoreactivity was almost always found in the nuclei of cancer cells (Figure 2). Positive cells were frequent in the advancing margin of the tumor. The standard morphologic criteria for identifing apoptosis were the presence of beaded or shrunken chromatin and apoptotic bodies with clear halos22). Almost all the positively stained cells and bodies were considered to be apoptotic cells that corresponded morphologically to the standard criteria of apoptotic cells (Figure 3). Nonspecific staining in necrotic foci showed faint, diffuse staining and this could be distinguished from the apoptotic nuclei by simple morphological examination.
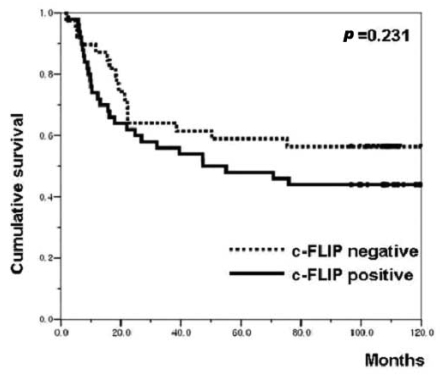
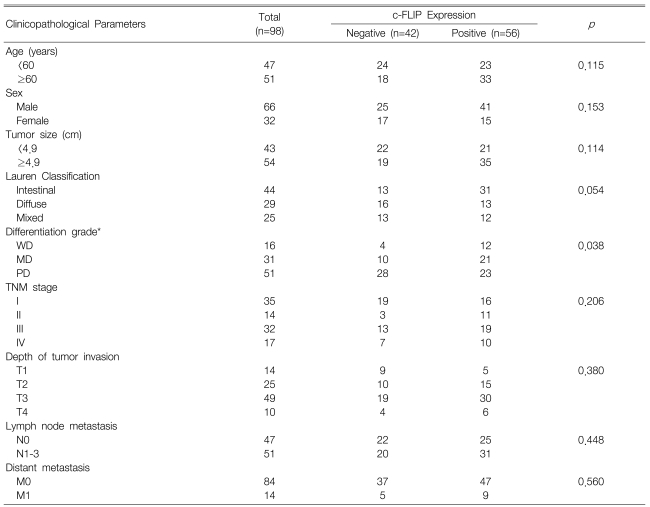
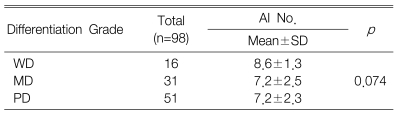
The correlations between the c-FLIP expression and the clinicopathological parameters are shown in Table 1. The expression of c-FLIP was significantly associated with histologic differentiation (p=0.038). However, there was no association between the c-FLIP expression and the depth of invasion, tumor stage, the status of the lymph nodes anddistant metastasis. Furthermore, the c-FLIP expression was not associated with patient survival (p=0.231) (Figure 4). When the KI, AI and the c-FLIP status and the clinicopathological parameters were analyzed by the Cox regression model, the KI, AI and c-FLIP status were not found to be independent prognostic factors (data not shown).
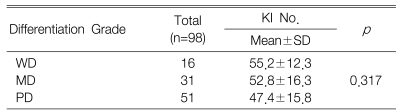
The KI for 98 tumors ranged from 7.6 to 85.0 with a mean KI of 50.4┬▒15.7. The mean KI value of the c-FLIP positive tumors was 54.1┬▒15.3 and this was significantly higher than that of the c-FLIP negative tumors (p=0.005) (Table 2). However, there was no difference between the differentiation grade and the KI (p=0.317) (Table 3).
Apoptosis is a cellular response that regulates important processes such as tissue homoeostasis, defense against certain pathogens and elimination of unwanted cells. Disturbance of this process by aberrantly extending the viability of cells or favoring an accumulation of transforming mutations is thought to contribute to carcinogenesis1-3). Apoptosis is a highly regulated process, and various inhibitors of this process are known to interfere with many steps of the extrinsic or the intrinsic pathway3, 4).
The expression of c-FLIP, which is a member the family of inhibitors of apoptosis, has recently been reported in various types of cancer, including gastric cancer9-13). Our study showed that the expression of c-FLIP is increased in gastric cancer tissue compared with the detectable levels in the matched normal gastric mucosa. These results suggest that the c-FLIPexpression is implicated in gastric carcinogenesis.
The c-FLIP expression has been reported to be associated with tumor progression9-13). However, in our study, there was no association between the c-FLIP expression and the depth of invasion, tumor stage, the status of the lymph nodes, distant metastasis or poor survival. Only the c-FLIP expression was significantly associated with the histologic differentiation. Previous studies on gastric cancer have reported contrary results. Zhou et al reported that the c-FLIP expression is associated with lymph node metastasis13), and Lee et al reported that there is no correlation between the histologic types and the c-FLIP expression12). These discrepancies may have cone about because of the different methods used to detect c-FLIP, the different scoring systems and the different antibodies used in immunohistochemistry as quantitative markers. At the protein level, there are two splicing variants for c-FLIP. They are the long splice form of c-FLIP (c-FLIPL) and the short splice form of c-FLIP (c-FLIPS). c-FLIPS contains only two N-terminal death effector domains (DEDs) that are very similar to the prodomains of caspase-8/-10. In contrast, c-FLIPL is identical in length with caspase-8, which also contains N-terminal tandem DEDs, but its caspase domain is altered, rendering it enzymically inactive. Therefore, c-FLIPL has the classical structure of a dominant-negative inhibitor23-25). Zhou et al used anti-c-FLIPL/S antibody13) and Lee et al used anti-c-FLIPL in their immunohistochemistry, the same as ours12). To date, since the exact roles of the two variants of c-FLIP in the extrinsic apoptotic pathway are not fully known, so further molecular and biochemical understanding of c-FLIP is necessary before any changes of this molecule can be applied in clinical practice as a prognostic factor.
Cell proliferation and apoptosis must be properly balanced in order to maintain tissue homeostasis1-3). However, tumor development and progression is generally regarded as dependent on an increased proliferation rate and an apoptosis rate that's too low to balance cell growth. The KI has been used to evaluate cell proliferation activity14, 15). In our study, the mean KI value of the c-FLIP positive tumors was significantly higher than that of the c-FLIP negative tumors. Chen et al reported that c-FLIP protein in endometrial carcinoma was significantly associated with the proliferating cell nuclear antigen-labeling index, which is known as another proliferation marker9). These results suggest that the c-FLIP expression may be associated with tumor cell proliferation.
c-FLIP is a recently identified antiapoptotic gene5-8). However, in our study, no significant difference was seen between the c-FLIP expression and the AI. There are several possible explanations for this contrary result. First, the steps of apoptosis are not dependent on c-FLIP alone, and these steps are regulated by many regulators, including survivin, Fas and nuclear factor-kappa B3). Second, in human cancers, the overall expression of c-FLIP as reported in different studies is difficult to compare due to the different score systems and the different antibodies the studies used. Third, the two splicing variants of c-FLIP may acts as an inhibitor or a promotor of the extrinsic apoptotic pathway7).Therefore, the exact biochemical understanding of the various inhibitors or promotors of the apoptotic pathway may provide the means to design drugs to correct the imbalance between apoptosis and proliferation, and this imbalance that is found in human cancers. A larger study would probably resolve this issue.
In summary, the expression of c-FLIP was increased in the gastric cancer tissues compared with the matched normal gastric mucosa. The expression of c-FLIP was significantly associated with histologic differentiation. However, there was no association between the c-FLIP expression and the depth of invasion, tumor stage, the status of the lymph nodes, distant metastasis and patient survival. The mean KI value of the c-FLIP positive tumors was significantly higher than that of the c-FLIP negative tumors. There was no significant difference between the c-FLIP expression and the AI. These results suggest that the c-FLIP expression may be associated with the tumor cell proliferation of gastric cancer.
References
1. Thompson CB. Apoptosis in the pathogenesis and treatment of disease. Science 1995. 267:1456ŌĆō1462PMID : 7878464.


3. Kiechle FL, Zhang X. Apoptosis: biochemical aspects and clinical implications. Clin Chim Acta 2002. 326:27ŌĆō45PMID : 12417095.


4. Salvesen GS, Dixit VM. Caspase activation: the induced-proximity model. Proc Natl Acad Sci U S A 1999. 96:10964ŌĆō10967PMID : 10500109.



5. Tschopp J, Irmler M, Thome M. Inhibition of fas death signals by FLIPs. Curr Opin Immunol 1998. 10:552ŌĆō558PMID : 9794838.


6. Yeh WC, Itie A, Elia AJ, Ng M, Shu HB, Wakeham A, Mirtsos C, Suzuki N, Bonnard M, Goeddel DV, Mak TW. Requirement for Casper (c-FLIP) in regulation of death receptor-induced apoptosis and embryonic development. Immunity 2000. 12:633ŌĆō642PMID : 10894163.


8. Wajant H. Targeting the FLICE inhibitory protein (FLIP) in cancer therapy. Mol Interv 2003. 3:124ŌĆō127PMID : 14993418.


9. Chen HX, Liu YJ, Zhou XD, Luo RY. Expression of cellular FLICE/caspase-8 inhibitory protein is associated with malignant potential in endometrial carcinoma. Int J Gynecol Cancer 2005. 15:663ŌĆō670PMID : 16014121.


10. Zhou XD, Yu JP, Chen HX, Yu HG, Luo HS. Expression of cellular FLICE-inhibitory protein and its association with p53 mutation in colon cancer. World J Gastroenterol 2005. 11:2482ŌĆō2485PMID : 15832422.



11. Korkolopoulou P, Goudopoulou A, Voutsinas G, Thomas-Tsagli E, Kapralos P, Patsouris E, Saetta AA. c-FLIP expression in bladder urothelial carcinomas: its role in resistance to Fas-mediated apoptosis and clinicopathologic correlations. Urology 2004. 63:1198ŌĆō1204PMID : 15183989.


12. Lee SH, Kim HS, Kim SY, Lee YS, Park WS, Kim SH, Lee JY, Yoo NJ. Increased expression of FLIP, an inhibitor of Fas-mediated apoptosis, in stomach cancer. APMIS 2003. 111:309ŌĆō314PMID : 12716387.


13. Zhou XD, Yu JP, Liu J, Luo HS, Chen HX, Yu HG. Overexpression of cellular FLICE-inhibitory protein (FLIP) in gastric adenocarcinoma. Clin Sci 2004. 106:397ŌĆō405PMID : 14636156.


14. McCormick D, Chong H, Hobbs C, Datta C, Hall PA. Detection of the Ki-67 antigen in fixed and wax-embedded sections with the monoclonal antibody MIB1. Histopathology 1993. 22:355ŌĆō360PMID : 8514278.


15. Weidner N, Moore DH 2nd, Vartanian R. Correlation of Ki-67 antigen expression with mitotic figure index and tumor grade in breast carcinomas using the novel "paraffin"-reactive MIB1 antibody. Hum Pathol 1994. 25:337ŌĆō342PMID : 8163266.


16. Chan AO, Luk JM, Hui WM, Lam SK. Molecular biology of gastric carcinoma: from laboratory to bedside. J Gastroenterol Hepatol 1999. 14:1150ŌĆō1160PMID : 10634150.


17. Lauren P. The two histologic main types of gastric carcinoma. Acta Pathol Microbiol Scand 1965. 64:31ŌĆō49PMID : 14320675.


18. Watanabe H, Jass JR, Sobin LH. Histologic typing of oesophageal and gastric Tumors. WHO International Histologic Classification of Tumors 1990. Berlin, Heidelberg, New York, Paris, Tokyo, Hong Kong: Springer-Verlag.
19. American Joint Committee on Cancer. Manual for staging cancer. Stomach cancer 1997. Philadelphia: Lippincott-Raven, 71ŌĆō76.
20. Reed JA, Manahan LJ, Park CS, Brigati DJ. Complete one-hour immunohistochemistry based on capillary action. Biotechniques 1992. 13:434ŌĆō443PMID : 1389176.

21. Gavrieli Y, Sherman Y, Ben-Sasson SA. Identification of programmed cell death in situ via specific labeling of nuclear DNA fragmentation. J Cell Biol 1992. 119:493ŌĆō501PMID : 1400587.



22. Wyllie AH, Morris RG, Smith AL, Dunlop D. Chromatin cleavage in apoptosis: association with condensed chromatin morphology and dependence on macromolecular synthesis. J Pathol 1984. 142:67ŌĆō77PMID : 6422024.


23. Chang DW, Xing Z, Pan Y, Algeciras-Schimnich A, Barnhart BC, Yaish-Ohad S, Peter ME, Yang X. c-FLIPL is a dual function regulator for caspase-8 activation and CD95-mediated apoptosis. EMBO J 2002. 21:3704ŌĆō3714PMID : 12110583.



Figure┬Ā1
Immunohistochemical staining of c-FLIP in the normal gastric mucosa (A) and the gastric cancer tissue (B). c-FLIP immunoreactivity was more intense in the cytoplasm of the cancer cells than that of the normal gastric mucosa (├Ś200).
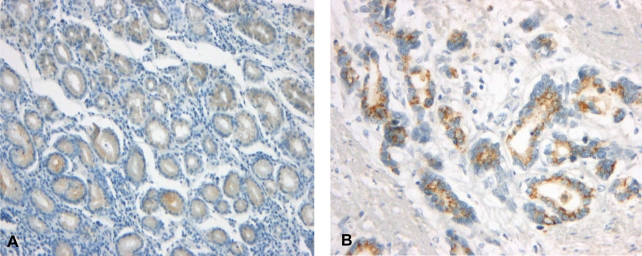
Figure┬Ā2
Immunohistochemical staining of Ki-67 in the gastric cancer tissue. Ki-67 immunoreactivity was found in the nuclei of the cancer cells (├Ś200).
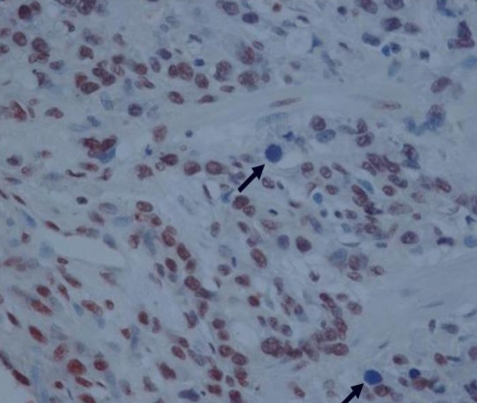
Figure┬Ā3
Detection of apoptotic cells and bodies (arrow) by TUNEL staining. An apoptotic body is characterized by a pyknotic nucleus surrounded by a shrunken cytoplasm, and the apoptotic body is separated from the surrounding cells by a halo (├Ś400).
Figure┬Ā4
Kaplan-Meier survival curve correlating disease specific survival with the positive (solid line) or negative (dotted line) expression of c-FLIP.
Table┬Ā1
Correlation between the c-FLIP expression and the clinicopathological parameters of gastric cancer



 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Download Citation
Download Citation Print
Print